Final report for GS23-278
Project Information
Recently, strawberry growers in Florida have incurred severe yield losses due to unprecedented outbreaks of leaf spot and fruit rot caused by a new species of Neopestalotiopsis. The failure of most fungicides registered for strawberry and the susceptibility of current cultivars demand breeding for improved genetic resistance. Fortunately, the University of Florida strawberry breeding program has identified a source of resistance to Neopestalotiopsis sp., an Israeli cultivar named ‘Yasmin’, and confirmed that resistance is heritable in breeding populations derived from ‘Yasmin’. We propose screening Yasmin-derived populations for resistance in inoculated field trials. Analysis of field resistance and genetic data via association mapping will approximate the locations of genes underlying resistance. Mining these locations for DNA diagnostic information in future work would enable breeders to effectively select for Neopestalotiopsis sp. resistance in their breeding populations. This research will support ongoing efforts to efficiently and more rapidly deliver cultivars with improved Neopestalotiopsis sp. resistance.
The overall aim of this project is to uncover genetic markers conferring resistance to Neopestalotiopsis sp. as targets for breeding resistant cultivars.
Objective 1: Evaluate Yasmin-derived populations for Neopestalotiopsis sp. resistance in replicated, inoculated field trials.
Objective 2: Identify genetic regions increasing resistance to Neopestalotiopsis sp. and characterize their phenotypic effects in Yasmin-derived populations.
Objective 3: Select individuals with superior resistance and other breeding-relevant traits for further backcrossing into elite backgrounds.
Cooperators
Research
 1. Sixteen full-sib families, generated from biparental crosses between Yasmin or progeny of Yasmin and 12 UF advanced selections with excellent yield and fruit quality, are retained as mother plants in the UF summer breeding nursery near Malin, Oregon. To develop large field populations, full-sib families, parents, and susceptible checks were clonally propagated via runners in the nursery. Five clonal replicates of each seedling were then harvested during mid-September and transported to the UF/IFAS Gulf Coast Research and Education Center (GCREC) in Wimauma, Florida where they were stored at 3.5°C until planting. Transplanting took place on 19 Oct. 2023 in a randomized complete block design with one runner plant in each of five blocks at GCREC. Each block consisted of a single raised bed (350 feet long and 3 feet wide) covered with plastic mulch. Plants were established with overhead irrigation during daylight hours for 10 days after planting, followed by drip irrigation for the rest of the growing reason. Once the seedlings were established, they were inoculated with an atomizer by spraying each plant with 2 ml of spore suspension (2 x 104 spores/ml). Spore suspensions were prepared from 7- to 10-day-old colonies of four Neopestalotiopsis sp. isolates revived from the culture collection at GCREC on potato dextrose agar under continuous fluorescent lighting at room temperature. After inoculation, plants were overhead irrigated for 10 days to encourage spore dispersal. Disease severity was assessed at four timepoints post inoculation based on the following 0-7 rating scale: 0, no symptoms; 1 = 1 to 10%, 2 = 11 to 25%, 3 = 26 to 50%, 4 = 51 to 80%, 5 = 81-100% of overall leaf area (per plant) covered by lesions, and 6 = dead. Phenotyping was initiated with the onset of symptoms on check cultivars and continued on a weekly or biweekly basis for four weeks post inoculation. A single area under disease progress stairs (AUDPS) value per individual was calculated by considering the severity score of all clonal replicates at all timepoints.
1. Sixteen full-sib families, generated from biparental crosses between Yasmin or progeny of Yasmin and 12 UF advanced selections with excellent yield and fruit quality, are retained as mother plants in the UF summer breeding nursery near Malin, Oregon. To develop large field populations, full-sib families, parents, and susceptible checks were clonally propagated via runners in the nursery. Five clonal replicates of each seedling were then harvested during mid-September and transported to the UF/IFAS Gulf Coast Research and Education Center (GCREC) in Wimauma, Florida where they were stored at 3.5°C until planting. Transplanting took place on 19 Oct. 2023 in a randomized complete block design with one runner plant in each of five blocks at GCREC. Each block consisted of a single raised bed (350 feet long and 3 feet wide) covered with plastic mulch. Plants were established with overhead irrigation during daylight hours for 10 days after planting, followed by drip irrigation for the rest of the growing reason. Once the seedlings were established, they were inoculated with an atomizer by spraying each plant with 2 ml of spore suspension (2 x 104 spores/ml). Spore suspensions were prepared from 7- to 10-day-old colonies of four Neopestalotiopsis sp. isolates revived from the culture collection at GCREC on potato dextrose agar under continuous fluorescent lighting at room temperature. After inoculation, plants were overhead irrigated for 10 days to encourage spore dispersal. Disease severity was assessed at four timepoints post inoculation based on the following 0-7 rating scale: 0, no symptoms; 1 = 1 to 10%, 2 = 11 to 25%, 3 = 26 to 50%, 4 = 51 to 80%, 5 = 81-100% of overall leaf area (per plant) covered by lesions, and 6 = dead. Phenotyping was initiated with the onset of symptoms on check cultivars and continued on a weekly or biweekly basis for four weeks post inoculation. A single area under disease progress stairs (AUDPS) value per individual was calculated by considering the severity score of all clonal replicates at all timepoints.
2. To detect DNA variants controlling Neopestalotiopsis sp. resistance, we analyzed all segregating populations via genome-wide association mapping (GWAS). A GWAS requires two data sources: DNA sequence information and seedling phenotypes. To obtain the first, DNA was extracted from all seedlings according to Mangandi et al. (2017) and submitted for genotyping using the FanaSNP array through Affymetrix. The FanaSNP array screens each seedling’s DNA with 50,000 single-nucleotide polymorphisms (SNPs) commonly segregating in cultivated populations. SNP genotypes of each seedling were then reported for the 50,000 examined locations across the strawberry genome. Coupled with phenotypic data collected from our seedling trial, SNP genotypes were analyzed via GWAS leading to the discovery of genetic variants that associate with Yasmin-derived resistance. The genetic effects and heritability of associated markers were estimated in statistical models to inform the next steps of breeding for Neopestalotiopsis sp. resistance.
3. Distributions of disease severity values were plotted for the progeny populations and compared against the susceptible and resistant parents’ mean values. Individuals with superior Neopestalotiopsis sp. resistance and adequate flower production were selected as donors of resistance in future crosses with proven, elite breeding parents, to recover desirable yield and fruit quality traits in successive generations.
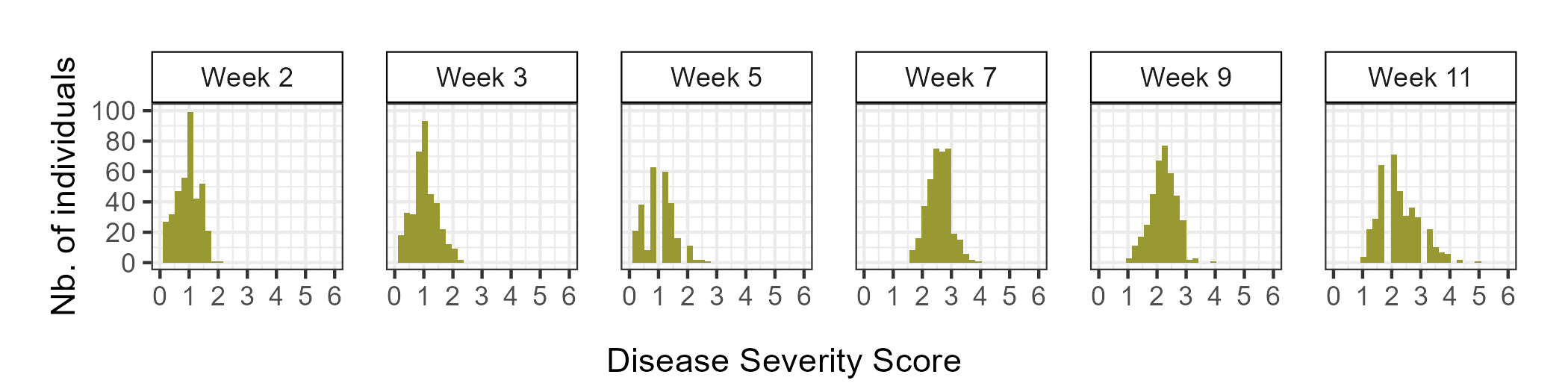
During the 2023-2024 strawberry production season, a total of 382 individuals were screened for resistance to Neopestalotiopsis sp. Individuals were categorized as either resistant (0 ≤ ȳ ≤ 2) or susceptible (2 < ȳ ≤ 6), with ȳ representing the mean of disease severity scores among replicates. Plants with a ȳ of 0 showed no symptoms, while those with a ȳ of 6 were dead. Out of the 382 individuals evaluated, 162 (42%) displayed mild symptoms (0 ≤ ȳ ≤ 2) and were classified as resistant, while 220 (58%) exhibited moderate to severe symptoms (2 < ȳ ≤ 6) and were classified as susceptible (Figure 1).
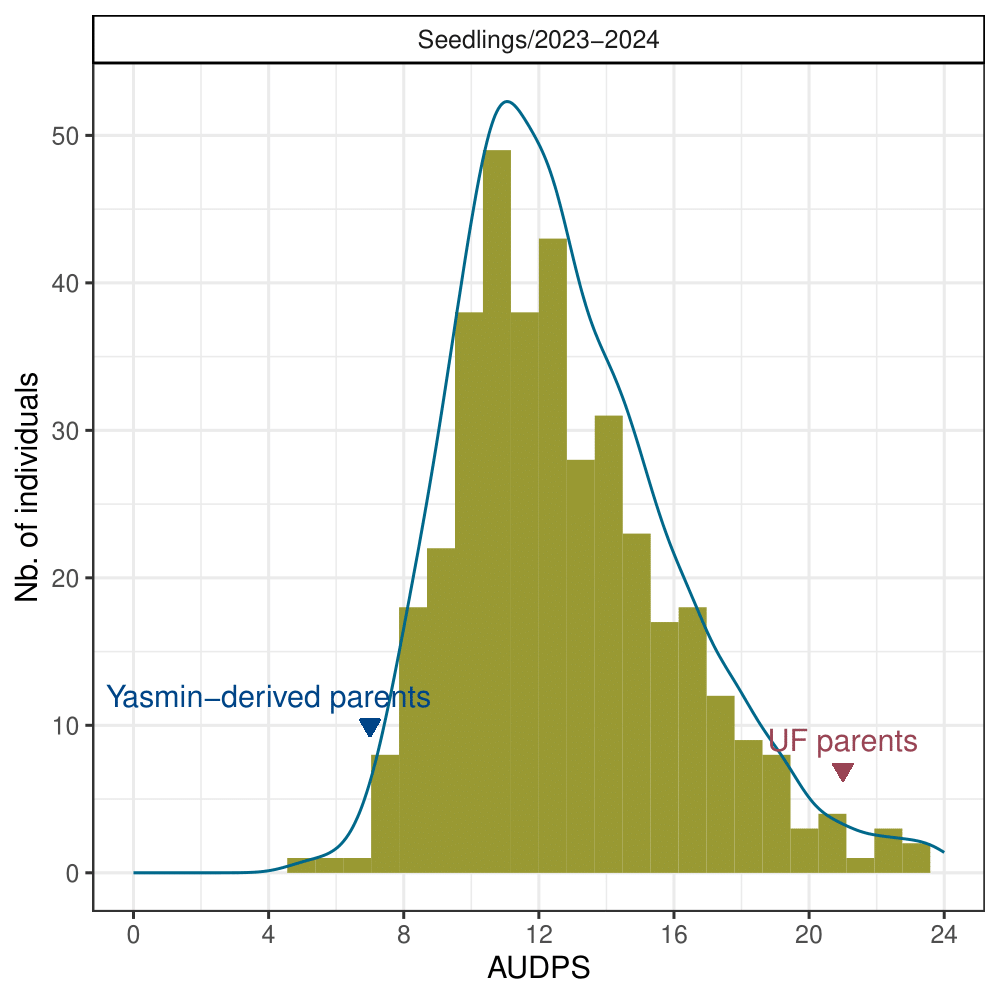
The mean values for the AUDPS revealed continuous distributions, skewed towards higher values indicative of increased disease severity due to wet conditions in Central Florida during the 2023-2024 production season (Figure 2). The heritability of resistance phenotypes was estimated at 0.69, indicating strong potential for breeding gains.
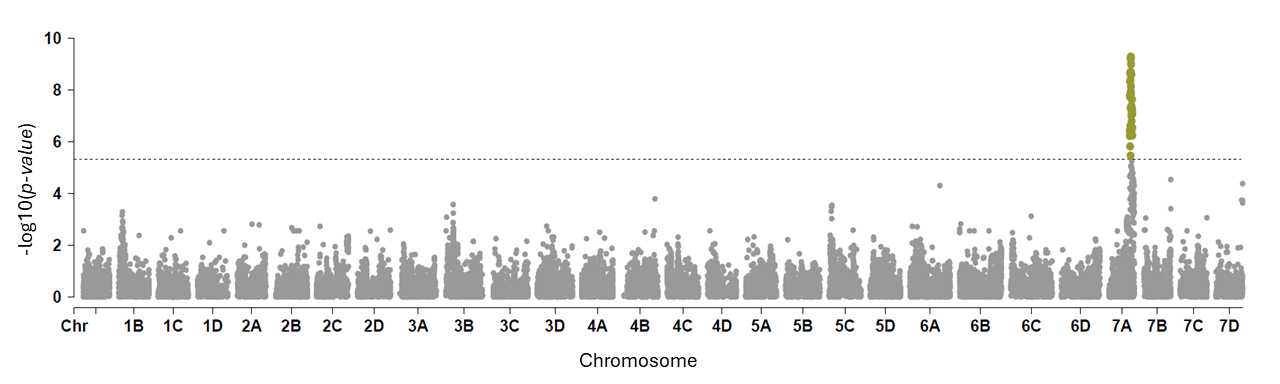
We analyzed the 382 individuals using a genome-wide association study (GWAS) with approximately 50,000 SNP markers to identify genetic associations with resistance to Neopestalotiopsis sp. GWAS revealed 20 significant SNP signals within a 700-kilobase region spanning 17.7–18.6 megabases on chromosome 7A of the strawberry genome. This locus, hereafter referred to as RNp2, is shown in Figure 3 (Manhattan plot), where the peaks indicate chromosome locations associated with increased resistance to Neopestalotiopsis sp.
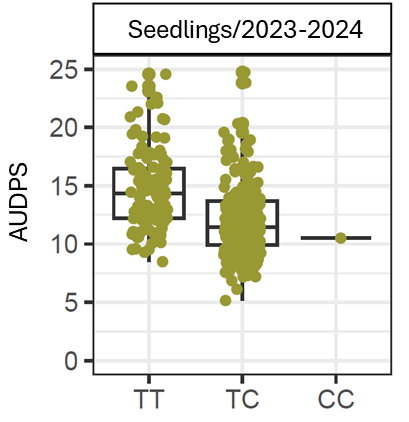 The most significant SNP within RNp2, AX-184266551 on the FanaSNP array, had a −log₁₀ p-value of 9.29. This marker has two alleles, T and C, with the C allele associated with greater resistance (Figure 4). AX-184266551 explained 13.8% of the phenotypic variance and reduced the mean AUDPS (Area Under the Disease Progress Stairs) score by 2.82 points per copy of the resistance allele (Figure 4). It correctly predicted resistance in 70% of the tested germplasm.
The most significant SNP within RNp2, AX-184266551 on the FanaSNP array, had a −log₁₀ p-value of 9.29. This marker has two alleles, T and C, with the C allele associated with greater resistance (Figure 4). AX-184266551 explained 13.8% of the phenotypic variance and reduced the mean AUDPS (Area Under the Disease Progress Stairs) score by 2.82 points per copy of the resistance allele (Figure 4). It correctly predicted resistance in 70% of the tested germplasm.
Overall, 42% of the progeny from the cultivar ‘Yasmin’ showed resistance, even under environmental conditions highly favorable for Neopestalotiopsis sp. outbreaks. Resistance appears to be partially controlled by the RNp2 locus on chromosome 7A. The introduction of RNp2 into University of Florida (UF) germplasm broadened the range of phenotypes toward resistance, with some plants showing no symptoms even 11 weeks after inoculation with the aggressive pathogen. In contrast, UF check cultivars developed severe leaf and fruit spots, and some plants had already died. These findings indicate that RNp2 is a strong candidate for marker-assisted selection (MAS). The significant SNPs identified here lay the groundwork for developing DNA-based diagnostic tools to help breeders efficiently select for Neopestalotiopsis sp. resistance in breeding populations. To our knowledge, this is the first report describing the genetic basis of resistance to Neopestalotiopsis sp. in strawberry.
In addition, we have identified promising individuals within Yasmin’s progeny that combine strong resistance to Neopestalotiopsis sp. with other desirable traits, such as high yield and fruit quality. These individuals will be immediately incorporated into the breeding program as parents in future crosses.
Educational & Outreach Activities
Participation summary:
We presented the roadmap for this research in a talk entitled “Recent Advances in Breeding for Neopestalotiopsis species and Macrophomina phaseolina in Strawberry” at the North American Strawberry Symposium (NASS) in San Luis Obispo, California. The audience included strawberry growers, breeders, and industry professionals across the production-to-consumer chain. The presentation updated attendees on our plan to use host resistance as part of the strategy for managing the emerging Neopestalotiopsis species and was followed by questions and open discussions throughout the symposium.
The results of this work were also presented in Elissar's public defense, entitled “Genomic Characterization of Resistance to Macrophomina phaseolina and Neopestalotiopsis Species in Strawberry and Breeding Applications,” at the Gulf Coast Research and Education Center in Wimauma, Florida. The audience included researchers, plant breeders, farm workers, and industry professionals. The defense concluded with questions and a discussion on future steps in Neopestalotiopsis sp. research.
In 2024, our findings were published in the The Plant Genome, a leading journal in the field, with acknowledgment of SARE funding. The publication has already been cited seven times and is available at https://doi.org/10.1002/tpg2.20477. Citation: Alam, E., Moyer, C., Verma, S., Peres, N. A., & Whitaker, V. M. (2024). Exploring the genetic basis of resistance to Neopestalotiopsis species in strawberry. The Plant Genome, 17(2), e20477.
Project Outcomes
During the course of this project, Neopestalotiopsis sp. has spread across strawberry nurseries and production fields in the U.S. (e.g., Ohio, North Carolina, Georgia, Texas, Indiana, Pennsylvania, and Florida), often moving undetected in asymptomatic transplants and thriving in warm, wet conditions. Chemical control options offer only limited suppression, especially in environments conducive to disease outbreaks, making host resistance the most sustainable management strategy.
Our research identified a strong, heritable source of genetic resistance to Neopestalotiopsis sp. in strawberry, along with DNA markers that enable breeders to target this resistance through marker-assisted selection. The resulting economic benefits include reducing disease-related yield losses and lowering pesticide application costs. This will help maintain the profitability of the Florida strawberry industry, currently dominated by susceptible cultivars, as well as other regions where Neopestalotiopsis sp. is becoming an annual concern. Socially, resistant cultivars will give growers more reliable harvests and greater resilience to disease outbreaks. Environmentally, replacing heavy chemical use with genetic resistance will reduce pesticide inputs, slow the development of pesticide resistance, and protect ecosystems, promoting long-term agricultural sustainability.
During the course of this project, we became even more convinced of the critical role host resistance plays in mitigating Neopestalotiopsis sp. outbreaks and sustaining strawberry production. Additional reports of susceptibility in elite germplasm, combined with increasing grower reports of outbreaks and evidence of the limited effectiveness of available fungicides under conditions favorable to disease, have reinforced the need to prioritize host resistance in management strategies. The risks of over-relying on fungicides effective against Neopestalotiopsis sp., such as Thiram and Switch, which are already widely used in rotations against Botrytis cinerea, have become clear with the increasing global incidence of fungicide-resistant Botrytis strains. All of this has strengthened our efforts to breed for durable genetic resistance to Neopestalotiopsis sp., not only to reduce chemical inputs and production costs, but also to protect long-term disease management tools.
Our results show that while ‘Yasmin’ is an excellent source of resistance, resistance to Neopestalotiopsis sp. in strawberry is likely quantitative, controlled by many loci of small to moderate effect. This means no single “silver bullet” gene provides complete protection. While screening diverse germplasm for additional resistance sources should continue, efforts should also focus on genomic selection, which enables the simultaneous selection of multiple loci. Our study populations, with publicly available phenotypic and genotypic data, provide a valuable resource for training and validating genomic selection models.