Progress report for GS24-315
Project Information
This project aims to evaluate the potential to develop, by plant breeding, Southern-adapted white lupin to fill both economic and environmental gaps. This project addresses the need for preventing winter soil erosion through cover cropping and identifying alternative plant-based protein sources for the Southeastern agriculture. White lupin (WL), a high N-fixing, P-acquiring, cool-season legume, holds unique potential to meet both goals. In the first year, we evaluated 200 genetically diverse WL lines from Auburn University’s breeding program and the USDA collection in a field trial targeting agronomic, root architectural, and physiological traits. Using Shovelomics and Rhizo imaging, we quantified 20 root traits and analyzed genetic variability through mixed models, PCA, correlation networks, and machine learning. Results revealed significant genetic variation for both aboveground and belowground traits, with strong heritability observed for dry biomass, maturity, and root features. Random forest models identified key root traits—such as RootPerimeter and Num_RootTips—as predictors of both grain yield and dry biomass, supporting the potential to breed for dual-purpose WL ideotypes.
To evaluate nitrogen fixation, whole-plant samples (stem, leaf, pod, and seed) from 400 plots have been ground and prepared for stable isotope analysis. Sample preparation for the second year of trials is ongoing, and essential materials (wells, tin capsules) have been acquired to support timely shipment and processing. For genomic analysis, leaf tissue will be collected in April 2025 and sent to the HudsonAlpha Institute for Biotechnology for genotyping. While genomic data is pending, UAV-based multispectral phenomic prediction has already shown promise, with R² = 0.30 for biomass and R² = 0.50 for grain yield depending on flight timing. Collectively, these advances lay the foundation for genomics- and phenomics-enabled development of white lupin cultivars tailored to the Southeast's agronomic and environmental needs.
2024-2025 Field trials are currently been phenotyped for all traits reported.
Objective 1: Assess WL diversity for genetic variability in traits related to cover crop and grain legume performance.
Objective 2: Explore the potential to improve N-fixation and P-acquisition by evaluation of root architecture and N-fixation ability.
Objective 3: Study the genetic diversity of WL and the prospect of using genomics to develop varieties rapidly.
Research
Germplasm Collection and Seed Increase Trials So Far:
From 2022 to 2023, we acquired 227 accessions of WL from the USDA NPGS. We also had 214 breeding lines unique to Auburn, derived from previous breeding efforts here. This includes an advanced WL selection we call “AU22” that is tall, but lodging resistant, and is alkaloid free, making it suitable for human/animal consumption. Our first trial with WL was primarily used to increase seed as we started with limited amounts per accession. We used this first trial and our in-progress 2023-2024 trial to characterize the germplasm relative for two traits: plant type and alkaloid-status. WL plants vary in their plant architecture, with some showing determinate growth (DET) and others indeterminate (INDET). Some, but not all WL plants genotypes sequester a potentially toxic alkaloid that requires post-processing before safe consumption. From our first trial, 337 genotypes produced seed: 196 were DET-type and were mostly from AU, 141 were INDET (mostly from USDA/NPGS lines). The DET types had no alkaloids, which makes them good for feed/food crops. AU22 and 28 USDA lines are the only indeterminate plant types without alkaloids.
In 2023-2024 we are conducting a preliminary yield trial with two-row 15-foot-long plots and two-replications each of the 200 lines selected from 2022-2023. We selected to test all 141 INDET types and the best of the DET types. This choice was to maximize diversity of INDET plant types while introducing more alkaloid-free genetics in our current trials and provides variation in maturity timing.
Objective 1: Assess WL diversity for genetic variability in traits related to cover crop and grain legume performance.
Experimental Design:
To achieve Objectives 1 and 2, we will conduct a two-year field evaluation for the project: Year 1 (2024-2025) and Year 2 (2025-2026). We will use the seed increase and data generated during the 2022-2023 and 2023-2024 seasons to select 200 lines from the available 441 WL accessions. We will use two 36”-spaced rows per 15-foot-long plot. Given sufficient seed is produced this year, we will test at two locations with two replications each. Trials will be conducted primarily at the EV Smith Plant Breeding Unit (PBU) in Tallassee, AL, and at a second site to be determined.
Data Collection: This objective focuses on agronomic data. All plots in both years will be regularly assessed for maturity timing using a weekly census. Stand count, growth in vegetation height/volume, and health indices will be captured using a multispectral-equipped drone, flown every 2-3 weeks. We will take notes on plant architecture (INDET vs. DET), alkaloid status, and flower color. Before seed harvest, biomass will be sampled (see Objective 2 below) as a cover crop performance indicator. At seed maturity, plots will be harvested by Combine for total seed yield and 100-seed weight.
Objective 2. Explore the potential to improve N-fixation and P-acquisition by evaluation of root architecture and N-fixation ability.
At maturity, but before seed harvest, we will harvest the above and belowground biomass from four plants per plot. Biomass samples will be dried at 60oC for 7 days and weighed. All aboveground samples will be ground finely to pass a 1 mm sieve and homogenized.
Carbon and nitrogen isotope analysis:
Between 3-5 mg of tissue from the PBU location will be weighed into tin capsules and submitted to the UC Davis Isotope Analysis Lab for assessment of C and N isotope content. The relative content of δ15N:δ14N is indicative of the proportion of tissue N derived from atmospheric N and thus from biological fixation (Sanz-Saez et al. 2019, Kalembasa et al., 2022). Furthermore, the ratio of C:N is critical for understanding the value of plant biomass both as a mulch (cover crop quality) and forage.
Analysis of cluster roots by imaging:
In both years, root systems of the same plants sampled for biomass and isotope analysis in field trials will be dug up and carefully cleaned. Roots will be dried, weighed and imaged to assess properties of their architecture, namely to detect variation in cluster root formation. The roots will be imaged using equipment operated by the AU Crop Physiology Lab. The platform (Fig. 2) plus RhizVision Explorer software can analyze images of roots taken from multiple angles simultaneously, to measure various traits, including length, diameter, volume, and branching patterns. This functionality makes it a valuable tool for investigating the root cluster and architecture of white lupin.
Data analysis: For Objectives 1 and 2, a mixed model will be used to assess the genetic and environmental variance components, as well as genetic parameters such as the heritability of the traits. WL lines will be ranked based on trait performance to identify the best-performing lines for each trait. Multivariate analysis will be used to understand and summarize the genetic correlations among traits, as these correlations determine how breeding improvements can / will proceed under selection. Cluster analysis will be used to classify the WL lines into distinct groups based on the similarity of their trait profiles. This classification can identify lines with desirable combinations of traits that can be targeted for selection in breeding programs.
Objective 3: Study the genetic diversity of WL and the prospect of using genomics to develop varieties rapidly.
We will use low-pass DNA-sequencing to obtain genome-wide single nucleotide polymorphism (SNPs) genotypes for the entire panel of WL. Genotyping the WL panel will unlock several possibilities. Firstly, to our knowledge, this will be the first genomic examination of the diversity of WL germplasm in North America. We will quantify genetic diversity, assess population structure and estimate genetic relatedness between and within the germplasm collections (NPGS and AU). Second, we will conduct genome-wide association analysis (GWAS) to identify genetic variants associated with traits-of-interest. This will be informative about the genetic complexity of the traits and, thus, the selection strategies that will be most effective. Finally, we will assess the prospect to use genome-wide selection (GS) by testing genomic prediction accuracy using cross-validation.
Materials and Methods:
Two hundred lines from the National Plant Germplasm System and Auburn University’s Cover Crop Breeding Program were evaluated for agronomic and root traits. Using Shovelomics and Rhizo imaging, we extracted 20 root traits (Table 1). Genetic variance, BLUPs, and heritability were estimated with ASReml-R. BLUP-based PCA, correlation heatmaps, and Machine learning Random Forest (ML-RF) identified key root traits affecting yield.

Objective 1: Assess WL diversity for genetic variability in traits related to cover crop and grain legume performance.
Results from linear mixed models show significant genetic variation among the white lupin lines tested (p>0.05) for flowering and pod maturity, plant height, grain yield, 100 seed weight, dry above-ground biomass, alkaloid content, and root architecture traits. This result was supported by PCA biplots (Figure 1) separated by plant type: Indeterminate vs. Determinate. It reveals genetic variability in trait expression across principal components (PCs), with clear clustering and vector dispersion between different plant types. RootTips, MaxRootWidth, RootDiameter, ConvexArea, and NetworkArea—cluster prominently in the right quadrants with strong projection along PC1 in both biplots. These traits exhibit high loading scores, indicating they are principal contributors to variation. Correlation heatmaps also show distinct trait correlation patterns between determinate and indeterminate types, highlighting trait architecture relevant to both cover crop (e.g., biomass, root traits) and grain legume (e.g., yield, maturity) performance. Strong positive correlations (Figure 2) among root morphological traits and between yield components and maturity traits indicate opportunities for multi-trait selection tailored to contrasting growth habits. Low to moderate broad-sense heritability was obtained for root architecture traits and grain yield, while dry biomass and maturity traits had moderate to high heritability (Figure 3).


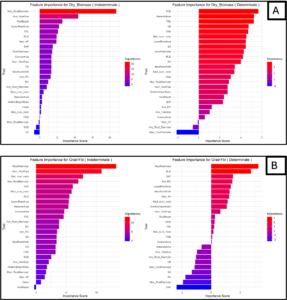
The random forest machine learning model (Figure 4 A and B) demonstrates that root traits are strongly linked to both cover crop performance (dry biomass) and grain yield in white lupin, with distinct trait importance profiles for Determinate and Indeterminate types. Key predictors such as RootPerimeter, Num_RootTips, and Max_RootDiameter consistently rank high, supporting the premise that optimizing root architecture can simultaneously enhance both biomass accumulation and reproductive output in a growth habit-specific manner.
Objective 2: Explore the potential to improve N-fixation and P-acquisition by evaluation of root architecture and N-fixation ability.
To evaluate N-fixation potential as outlined in the proposal, we have completed tissue grinding for whole-plant samples (stem, leaf, pod, and seed) from 400 plots of the 2023–2024 field trial. Sample preparation for the 2024–2025 field trial is currently underway and will follow the same protocol. All samples will be submitted for stable isotope analysis to quantify biological nitrogen fixation. The required materials for sample processing and shipment, including wells, tin capsules, and other consumables, have already been purchased in preparation for analysis.
Objective 3: Study the genetic diversity of WL and the prospect of using genomics to develop varieties rapidly.
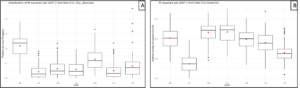
To evaluate the genetic diversity of white lupin (WL) and enable rapid varietal development, young, healthy leaf tissue will be collected from all accessions and shipped to the HudsonAlpha Institute for Biotechnology for genotyping in April 2025. Genomic data generation is expected to take approximately three months, after which the first round of genomic prediction (GP) will be conducted in Fall 2025, targeting key traits such as grain yield and biomass. While genomic data is pending, phenomic prediction using multispectral UAV imagery has already demonstrated strong potential. Preliminary results achieved R² = 0.30 for dry biomass and R² = 0.50 for grain yield (Figure 5 A dry above-ground biomass and 5 B grain yield), with prediction accuracy varying by flight date. These findings underscore the value of UAV-based high-throughput phenotyping and support an integrative selection strategy combining phenomics and genomics to accelerate white lupin breeding.
Educational & Outreach Activities
Participation summary:
We have recently made significant progress analyzing data from last year's trial. Poster presentations, e.g. at the CSSA/ASA/SSA Conference are planned for 2025.
Several undergraduate researchers have been involved in the project. One of them is planning Msc research based on experience working on this project.
