Final report for GNE22-275
Project Information
Specialized metabolites act as a carbon source on leaf surfaces and have antimicrobial properties. However, specialized metabolites are a function of plant metabolism which is influenced by the plant stress response. Changes in specialized metabolite profiles during stress responses have already been linked to changes in microbiomes within the rhizosphere (plant roots) but this has not been explored in the phyllosphere (leaf surface). The purpose of this work was to examine the influence of altered metabolite expression via induced stress on phyllosphere bacterial assemblages. By pairing untargeted metabolomic techniques with cellular biology, this intended to tease apart the influence host stress response has on the leaf surface metabolic profile (LSMP) and the subsequent effect this has on leaf-associated bacteria. By focusing on the tomato-Pseudomonas syringae pv. tomato DC3000 (DC3K) model system, researchers had two objectives: 1) characterize how induced plant defense responses influence leaf surface metabolite profiles and 2) subsequently their impact on phyllosphere bacterial assemblages.
Characterizing changes in leaf surface metabolites was done by subjecting tomato plants to biotic stress via inoculation with DC3K. Three days after inoculation, metabolites were extracted from the infected leaf and distal to infected leaf (distal), and analyzed using nuclear magnetic resonance spectroscopy (NMR). The resultant hydrogen peaks in the NMR spectra were cleaned, aligned to EtOH as a standard, peak selection noise was minimized, and peaks were manually organized until all peaks were validated. These peaks were identified for significance, and the significantly different peaks were organized into a list and dereplicated into possible natural products. The most notable changes that occurred to LSMP during pathogen infection were high levels of unique phenolic compounds in both treated and distal leaves, and increased sugar abundance in leaves distal to pathogen infection.
Subsequently, understanding the impact of these changes was the second objective. This was done utilizing an in vitro and in vivo approach. For both approaches, a plant growth promoter was introduced, Pseudomonas fluorescens A506, to compare the effects of the pathogen DC3K to a closely related species.
The in vitro approach grew DC3K or A506 in M9 minimal media with no carbon sources and leaf surface metabolites extracted from either the treated or distal leaves from control plants or DC3K infected plants. The bacteria were grown in a 96 well plate to observe changes in OD over 18 hours. Analysis of these results indicated leaf surface metabolites extracted from distal leaves in pathogen stressed plants facilitated the growth of both DC3K and A506, suggesting a complex interplay between plant metabolism and microbial colonization.
The in vivo approach inoculated tomato plants with either DC3K or A506 to induce biotic stress. The inoculation was performed by suspending the respective bacterial species in a liquid suspension and dunking the designated leaves to reduce contamination likelihood. After 72 hours, the distal leaves were further inoculated with either DC3K or A506, depending on the experimental treatment group. This step allowed for the observation of bacterial establishment on subsequent leaves following the initial stress induction. Plants infected with the pathogen DC3K exhibited a significant increase in pathogen population establishment on subsequent leaves, up to threefold greater than populations established on uninfected plants. This finding indicates a systemic adjustment in host LSMP, potentially favoring pathogen establishment and persistence within the plant environment. Furthermore, the presence of A506 on treated leaves appeared to induce systemic susceptibility to DC3K, resulting in increased pathogen population establishment on subsequent leaves. However, A506 populations were not significantly affected when grown in DC3K-infected plants, suggesting a selective advantage conferred by altered LSMP in favoring pathogen establishment.
The study provides a comprehensive understanding of the intricate interplay between plant defense responses, leaf surface metabolite profiles (LSMP), and phyllosphere bacterial colonization dynamics. In time, altering the metabolic profiles of tomatoes may be a strong strategy to inhibit pathogen establishment. Furthermore, the use of plant growth promoters without sufficient evidence of their efficacy within a real agriculture system may worsen plant tolerance to pathogen infection. Understanding metabolic drivers or host mediated responses that lead to LSMP changes would minimize the need for toxic pesticides that drive pathogen resistance and reduce farmer profit.
Objective 1: Characterizing the influence of biotic stress on leaf surface metabolite profiles. Plant defense responses to pathogens are dynamic, multifaceted, and occur at multiple spatial scales. The well-established host-pathogen model system, tomato (Solanum lycopersicon) and Pseudomonas syringae pv. tomato DC3000 (DC3K; bacterial speck), will be used to investigate the relationship between induction of plant defense responses and its subsequent effects on leaf surface metabolite profiles (LSMP). Defining LSMP changes on leaves directly affected by the pathogen, as well as leaves that have not been treated but are on the same plant will give a better understanding of the influence of host stress response on the entire plant. Quantification of LSMP will be done utilizing nuclear magnetic resonance.
Object 2: Investigating biotic stress influence on phyllosphere bacterial assemblage establishment. Metabolites accessible to phylloplane colonizing bacteria influence assemblage diversity [16-18]. However, work has not characterized how induced plant defense responses, which directly alters the host metabolite expression, influence associated phyllosphere communities. To address this, bacterial establishment on the tomato phyllosphere post defense induction will be observed. This aim will resolve patterns of bacterial establishment on leaves of tomato that have been stressed with DC3K. Characterization of changes in colonization will be done by relating leaf surface metabolites changes induced by DC3K to changes in DC3K population size. This will be compared with the population size establishment of a closely related species Pseudomonas fluorescens A506, to compare differences in effect. Changes in species composition associated with patterns of defense elicitation and spatial distance from the initial site of infection will be examined via cfu counting.
The purpose of this project is to examine the influence of host defense, and subsequent altered metabolite expression, on phyllosphere (leaf surface) microbiomes. It is my central hypothesis that metabolite profiles on the leaf surface selectively influence phyllosphere colonization and establishment, and that these metabolite profiles are linked to metabolic changes in the host which are a trait that can be altered via breeding.
The use of conventional agricultural methods (e.g. pesticides) to increase crop yield has previously allowed generations to break through established yield plateaus . However, overuse of foliar pesticides jeopardizes the ability of future generations to meet their food needs due to increased gain of resistance in pathogens and runoff causing ecosystem disruptions. Alternative options to pesticides include foliar applications of biological control agents (BCA) and utilization of resistant cultivars, either with resistance genes or horizontal resistance breeding packages, to minimize disease. BCA have found success under highly controlled settings, but are limited in field applications. Typically this reduced effectiveness is attributed to climate events (e.g. rainfall) and competition with already established microbiomes. Alternatively, resistance genes are effective within fields but can be overcome by pathogens ability to circumvent these mechanisms. Horizontal breeding, or breeding of host traits that compound to reduce disease, only reduce disease severity. Therefore, none of these options provide long-term, sustainable solutions capable of stopping disease incidences. However, mixing the use of BCAs and horizontal resistance breeding, specifically for specialized metabolites, is an innovative solution for long term sustainable agriculture.
A significant limitation of BCA application in field is persistence but this can be overcome by expressed host metabolism on the leaf surface, a trait that can be bred. Studies have shown that specialized metabolites produced by plants such as essential oils, phenolics, and alkaloids influence the bacterial communities that establish on the leaf surface. These compounds can either have antimicrobial effects or they can be used as carbon sources by phyllosphere endophytes. Furthermore, research on other components of the plant (e.g. rhizosphere) has showcased the capacity for induced defense responses to alter expressed metabolites to select for microbiomes that contain beneficial organisms. Within the phyllosphere, studies have only looked at specialized metabolites on leaf surfaces at a single time point and do not identify potential shifts in these metabolites over time during stress responses. This work, therefore, aims to define the relationship between altered states of plant metabolism and its subsequent effect on phyllosphere assemblages. This work utilized an untargeted metabolomics approach, coupled with cellular biology, to study the effect stress induction has on the composition of phylloplane metabolites and subsequent bacterial assemblages. Altered metabolic states likely disrupt established microbiomes which can lead to increased BCA penetrance and persistence, especially if paired with metabolizable specialized metabolites. Altering the plant metabolism is possible through breeding, therefore pairing BCA and horizontal resistance can result in robust protection for plants by selective for beneficial microbiomes.
Research
General Experimental Procedures
Experiments were conducted in an indoor growth chamber at the Penn State University's Plant Pathology and Environmental Microbiology department. This growth chamber was a single, isolated, full environment Conviron chamber which allowed for robust control over all abiotic conditions. The utilized growth chamber contained two compartments, with equal lighting in the top growing compartment and the lower growing compartment. This was validated utilizing a light meter software app called 'photone'. Independent testing with comparisons to hardware light meter (Extech LT40 LED Light Meter) and Photone showcased near identical readings between attempts (~.5 +/-) and photone was utilized for the rest of the study. Table 1 below provides details on the growth chamber conditions, and seed sterilization protocols. In order to assure that plants had minimal disturbance by a nascent microbial community which could act as bio stimulants, mountain fresh plus seeds (MFP) were sterilized and grown in glass petri dishes for 1 week, typically until tap roots were ~ one (1) inch long. After a week, plantlets were transplanted into ~ five (5) inch pots with growing media. Labels were distributed randomly to each plant, and then each biological replicate per treatment was randomly selected and put into a tray with biological replicates of other treatments. This ensured all treatments were distributed equally across the top and bottom of the growth chamber and maintained randomization that could be included for later statistical analysis as a 'block'. There were in total five (5) biological replicates across five (5) 'blocks' and two (2) 'levels' (the top growing compartment or the lower growing compartment in the growth chamber). For both objectives in this experiment, plants were allowed to grow two weeks post transplanting before initial inoculations. The second youngest fully mature leaf for each plant was selected and considered the 'treated' leaf marked with a white band (Figure 1, red; https://projects.sare.org/?attachment_id=1072763) and the next youngest fully mature leaf was considered the 'distal' leaf (Figure 1, white; https://projects.sare.org/?attachment_id=1072763). Inoculations were done by suspending the bacterial species of interest into a liquid suspension of KB at one (1) OD (a measurement of bacterial mass in suspension, one (1) OD is ~ two (2) million cells) and submerging the 'treated leaf' in the suspension for twenty (20) seconds.
Table 1. Details on growth chamber conditions and seed sterilization protocols.
| Growth Chamber Conditions | Light: 16hr on, 8 hr off Relative humidity: 90% when not inoculated, 95% when inoculated |
| Seed sterilization protocol |
|
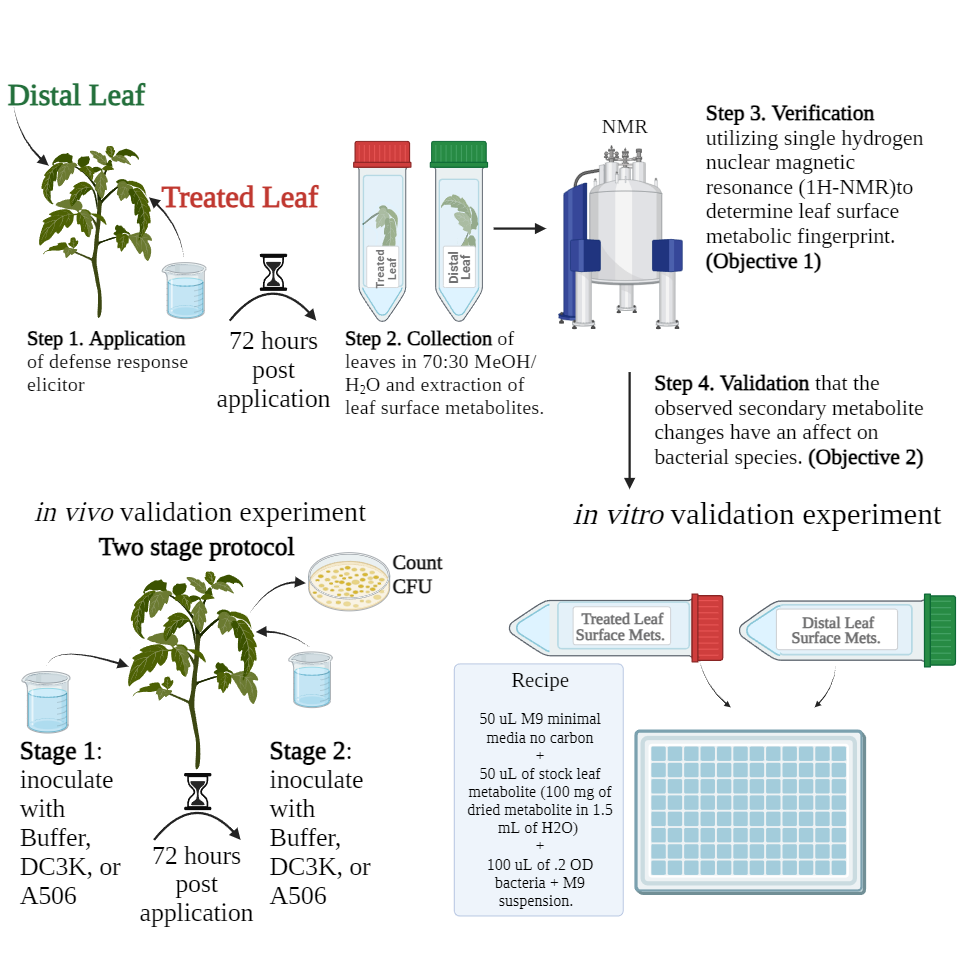
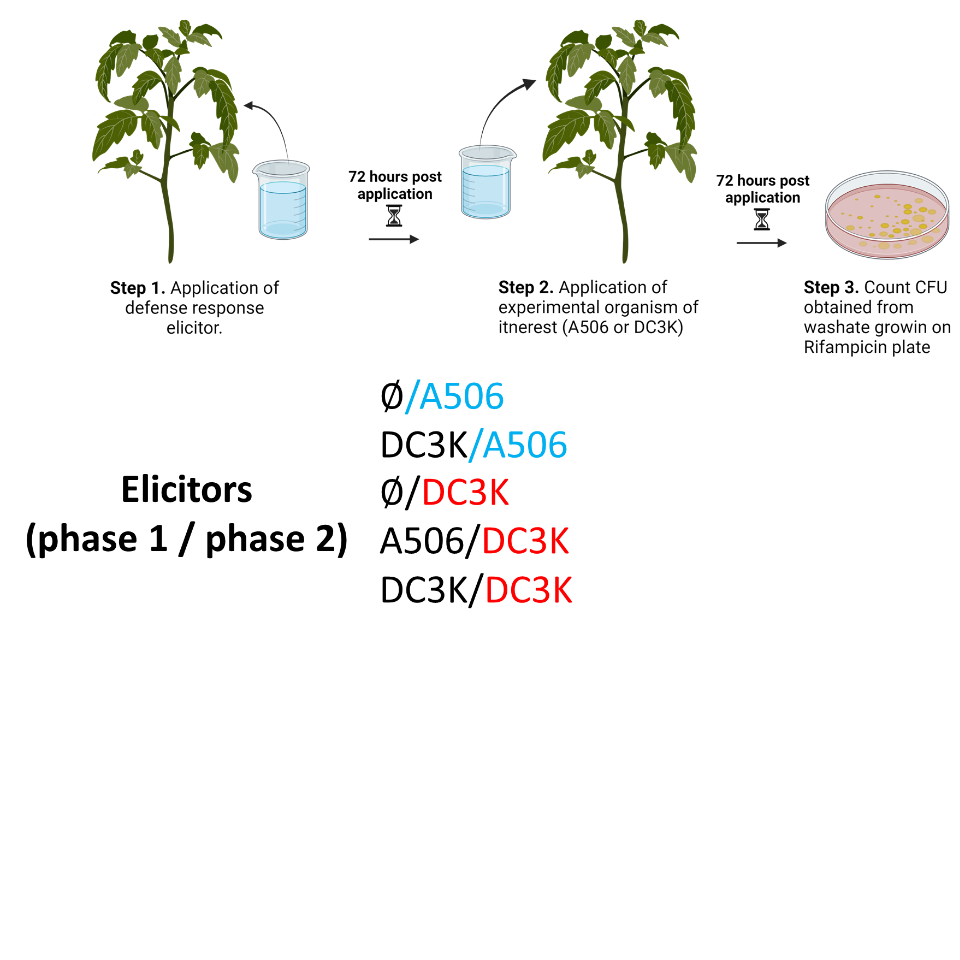
This work was conducted in two parts that subsequently filtered into each other (Figure 2 and Figure 3). Each experiment will be explained in further detail below. Experiments and their timelines, treatments and controls, and subsequent notes for both experiments can be found in table 2.
Table 2. Experiments and their timelines, treatments and controls, and subsequent notes
| Experiment | Timing | Treatments | Notes |
| Leaf Surface Metabolite Extraction | Started September 2022, metabolite extraction and HNMR dereplication finished July 2023 | Control treatment: application of 10 mM MgCl2 buffer + .001% silwet Pathogen treatment: application of 10 mM MgCl2 buffer + .001% silwet + 1 OD pathogen suspension | In this experiment, the hardest thing was getting metabolites dried down. Almost an entire year was spent fully optimizing the extraction protocol. |
| In vitro assessment | Started August 2023, ended October 2023 | control treatment negative: M9 + metabolites control treatment DC3K positive: M9 no carbon source + DC3KControl treatment A506 position: M9 no carbon source + A506 Pathogen treatment: DC3K M9 without carbon source + Secondary MetabolitesA506 treatment: A506 + M9 without carbon source + Secondary Metabolites | In this case, both controls were needed as assessments against DC3K and A506 could not be done. Assessments were done comparing controls of A506 to treated A506 and treated DC3K against control DC3K. Crosswise comparisons of growth between DC3K and A506 are challenging to do besides addressing trends. |
| Two stage elicitor protocol (in vivo) | Started November 2023, ended January 2024 |
Negative Control ∅/∅ (double negative, only buffer applied to each leaf) Control ∅/DC3K (buffer added to treated leaf, DC3K to distal leaf) Control ∅/A506 (buffer added to treated leaf, DC3K to distal leaf) DC3K/A506 (DC3K added to treated leaf, A506 to distal leaf) A506/DC3K (A506 added to treated leaf, DC3K to distal leaf) DC3K/DC3K (DC3K added to treated leaf,DC3Kto distal leaf) |
CFUs were counted on petri dishes with rifampicin. This allowed for only culturing either A506 or DC3K from the samples. This is why the double negative is not represented in subsequent graphs, there was no growth. This also indicates that the dipping technique minimized contamination. |
Experimental Procedure For Objective 1
The first portion of the experiment was to validate if leaf surface metabolites changed as a function biotic stress. Stress was induced by infecting the treated leaf with the pathogen Pseudomonas syringae pv. tomato DC3000 (DC3K), which is known to cause bacterial speck (Picture 1)

Seventy-two (72) hours after inoculation with the pathogen, the treated leaf and distal leaf were collected into separate 50 mL falcon tubes, submerged in MeOH/H2O (70%/30%), vortexed for ten (10) seconds, sonicated for ten (10) minutes, and vortexed again for ten (10) seconds before being being discarded. For each sample, the MeOH/H2O solution was then filtered through a .22 um PES single use filter (150 mL) and poured onto two petri dishes to dry (Figure 4).
All samples finished drying twenty-four (24) hours after pouring onto the petri dishes. One set of dried Metabolites were resuspended in .5 mL of deuterated H2O while the other half was frozen and saved for use in the second objective of this study. The reconstituted metabolites in .5 mL D2O were aliquoted into a 5 mm NMR tube. Single proton nuclear magnetic resonance imaging allows comparison of hydrogen composition across samples where differences in the position of the hydrogen on the spectrum indicates a different hydrogen (Figure 5).
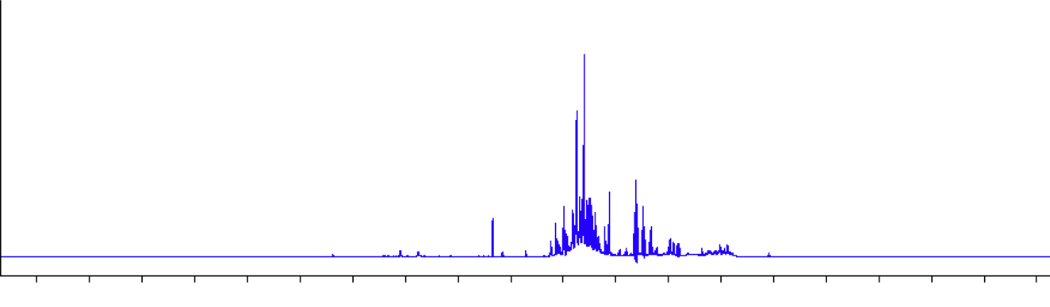
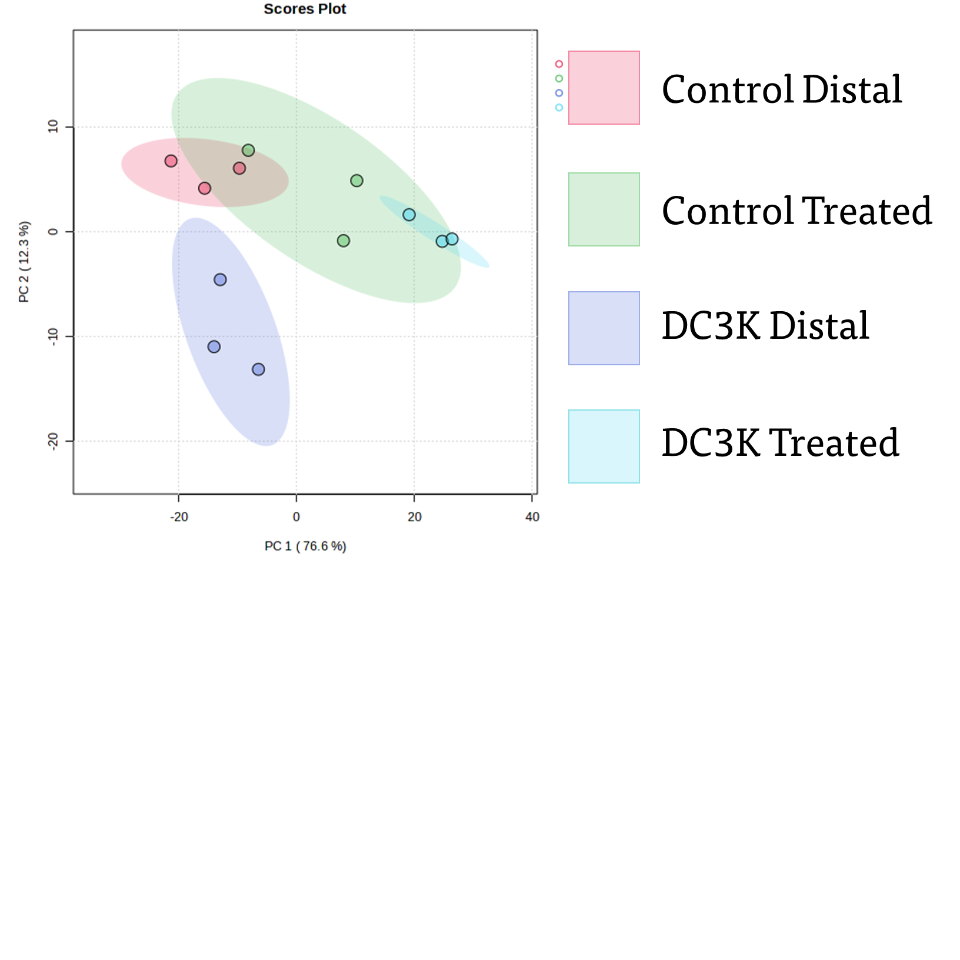
Notably, this created a 'fingerprint' capable of comparing changes in the metabolic composition between the control and pathogen treatment via differences in peak position and intensity. Generated spectra from HNMR on samples were aligned by EtOH peak position (ppm ~2.3, not shown above) and normalized by leaf weight (data not shown here). Spectra were then filtered by a noise value threshold of 100 to establish a 'floor' for relevant peaks. Peaks that persisted past this filtering step were then assigned as either 'overlapping', 'control associated', or 'DC3K associated' via manual alignment. Peaks that overlapped were given the same number and peaks that did not were given a unique number. For example, hydrogen peak at ppm 3.49 was consistent in all spectra regardless of sample and was assigned the character 1 to indicate character 1 relates to the ppm 3.49. At character 1, all samples had a peak at ppm 3.49. Peak 7.43 was only identified in pathogen samples and was given the character 2. No control samples had a peak at character 2, so each control sample had a 0 value, and each pathogen sample had a value of 7.43. So on and so forth, until all peaks had been given a character and that character was either considered to be an overlapping peak, a control associated peak, or a DC3K associated peak. A value of 0 was input for empty values where the peak was not present. This created a character by value data frame which allowed comparisons of peak composition without the expectation of them being additive values. Notably if not done this way, most statistical programs either 1) added the values to compare means which has no relationship to the experiment as these are discreet positions, or 2) assumed that the value was not present if not aligned within the same row. The aligned and binned peak values were input into metaboanalyst's one factor statistical analysis module and generated a PCA of identified peaks across control and pathogen plants, in both treated and distal leaves. These significantly different peaks were then input into a natural products dereplication process wherein the peaks are compared to known spectra to isolate series of peak combinations that can be identified within the fingerprint and resolved to a metabolite or other product.
Experimental Procedure For Objective 2
The second portion of this experiment was to understand how the changes in leaf surface metabolites affect the pathogen DC3K and a closely related species that is considered a plant growth promoter, Pseudomonas fluorescens A506 (A506). The first step for validation was an in vitro 96-well plate experiment observing changes in bacterial growth to understand if leaf surface metabolite changes observed in Objective 1 were related to metabolites with antibiotic capacities or if the altered metabolite profile promoted pathogen growth. The second step for validation was a two-stage in vitro experiment, wherein seventy-two hours after initial biotic elicitor inoculation (DC3K or A506) the distal leaf was inoculated with either DC3K and A506 and CFUs were recovered from distal leaves seventy-two hours post distal inoculation.
in vitro validation
DC3K and A506 were grown from single colony streaks for twenty-four 24 hours prior to application. On day of application, each species was suspended in M9 minimal media with no carbon source (recipe Table 3) to an OD of .2. The frozen metabolites collected from Objective 1 were reconstituted to 1.5 mL utilizing the same M9 no carbon media. 500 ul of M9-bacterial solution was placed in each well, and 500 ul of M9-metabolite solution were placed in each well. Growth of bacteria over 18 hours was recorded, and this experiment was repeated 3 times for a total N of 15 for each treatment/species.
| M9 Minimal media (no carbon) |
M9 Minimal Media salts minus nitrogen sourceFor 1L 5x M9 salts: 64g Na2HPO4-7H2O 15g KH2PO4 2.5g NaCl H2O fill to final 1L volume and autoclave To prepare 500mL M9 minimal media; 100mL of 5xM9 salts 1 mL 1M MgSO4 50 uL 1M CaCl2 2.5 mL filter sterilized NH4Cl autoclaved H2O to final volume of 500mL |
in vivo validation
Stress was induced by inoculating the treated leaf with either pathogen DC3K or growth promoter A506. Seventy-two (72) hours after inoculation with either bacteria, the distal leave was treated with either DC3K or A506. For controls, there was a fully negative control (inoculated distal leaf and treated leaf with buffer), a DC3K control (buffer treated leaf and DC3K inoculated distal leaf), and an A506 control (buffer treated leaf and A506 inoculated distal leaf). 72 hours after distal inoculation, only the distal leaves were collected across all treatments, and samples were placed into separate 50 mL falcon tubes filled with KB buffer. Submerged leaf samples were vortexed for ten (10) seconds, sonicated for ten (10) minutes, and vortexed again for ten (10) seconds before being being discarded. The resulting KB+CFUs suspension was diluted in 1/5 dilutions in subsequent KB for a total of 8 times and pipetted onto a rifampicin antibiotic plate. A506 and DC3K strains used in this experiment were rifampicin resistant. This procedure allowed for comparisons between what an initial infection would look like (control DC3K or control A506) and a subsequent infection on the distal leaves would look like, while maintaining equal time for the infection to initiate. This resulted in CFUs that could be corroborated back to the treatment for comparison and normalized by plant surface area.
Changes in leaf surface metabolite profiles due to pathogen stimulation
Pathogen infection causes changes in leaf surface metabolite profiles associated both at the site of infection and in the distal components of leaves (Figure 6).
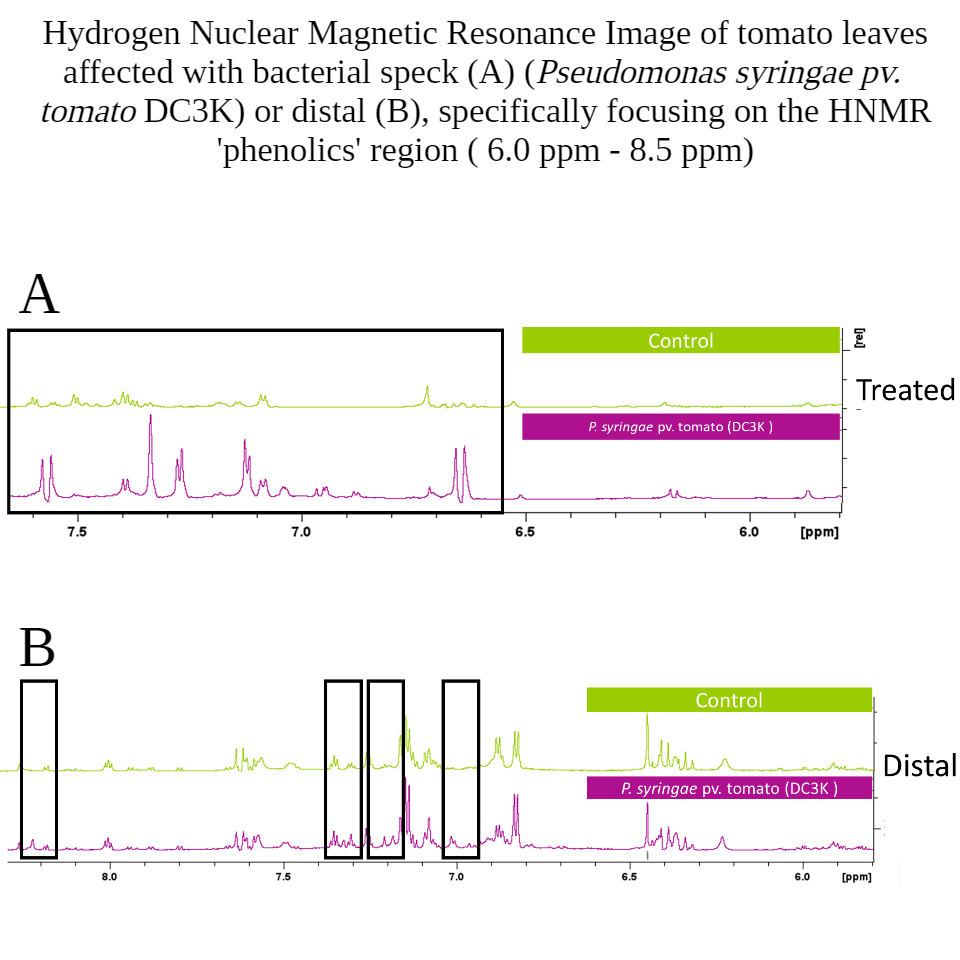
There were very few amino acids that were significantly enriched, and the three amino acids that were identified also shared dereplication with other secondary metabolites. The presence of novel aromatic phenolics in both treated and distal leaves was extremely fascinating, and corroborated the likelihood of manipulated Jasmonic Acid defense responses by the phytohormone mimic coronatine that is produced by DC3K; this is one explanation for the result of increased accumulation of secondary phenolic metabolites (Figure 7).
Furthermore, significant increases in the sugar profile of distal leaves infected by DC3K were identified (Figure 8),
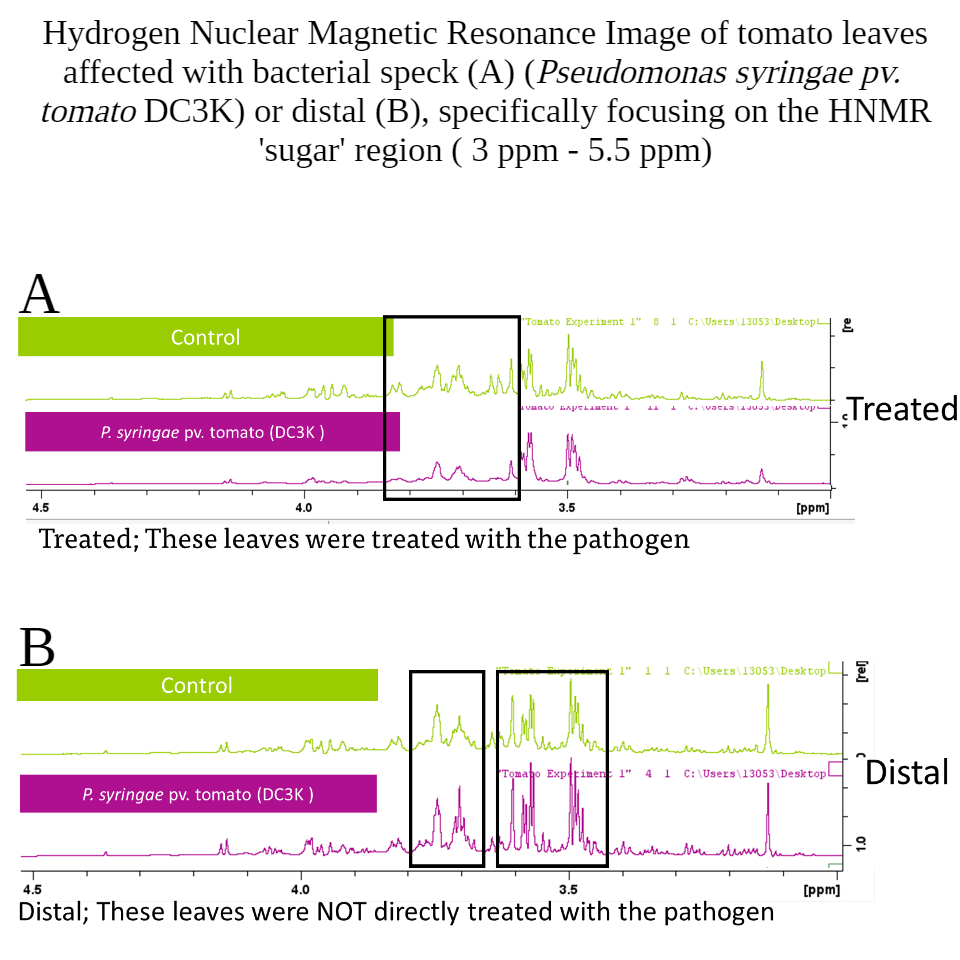
and significantly reduced sugar in the treated leaf aligns with the increased bacterial population presence. A note here, that a lot of metabolites are stripped from the leaf surface when dipping the leaf into the bacterial suspension (Figure 9), which may result in reduced sugars on the treated leaf surface.
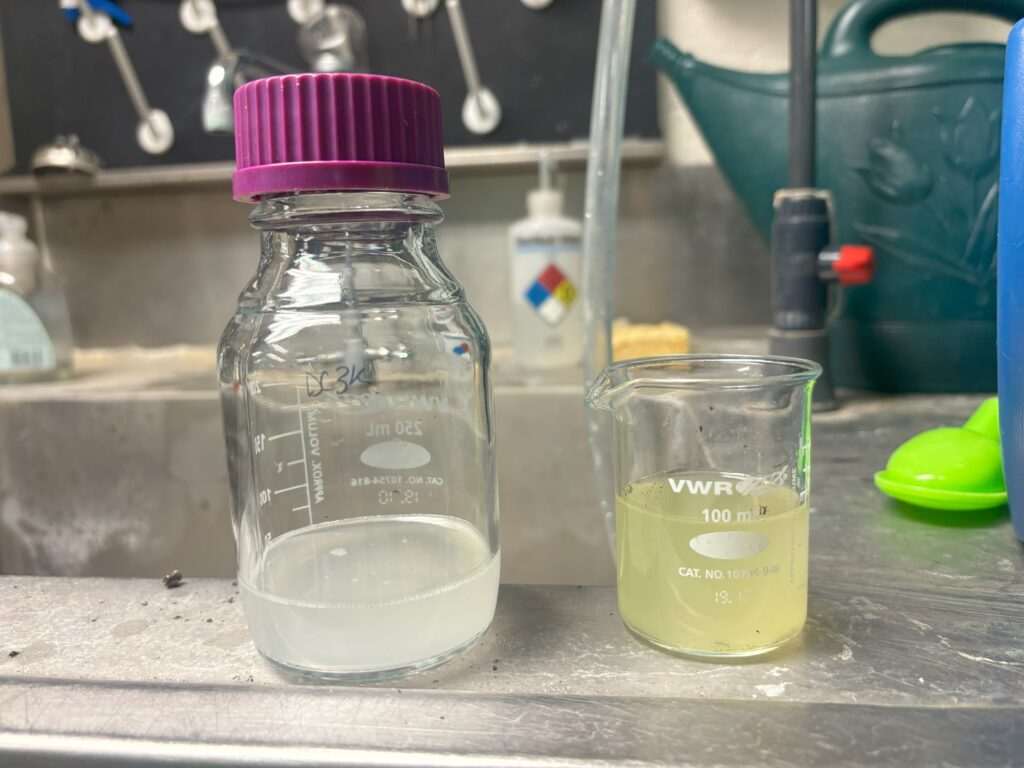

Taken together, a significant enrichment of phenolics occurs in tomato leaf surfaces during pathogen infection, both at the site of treatment and in an uninfected distal leaf. This provides evidence for a systemic adjustment in host leaf surface metabolites which has not been recorded before.
Effects of leaf surface metabolites in vitro
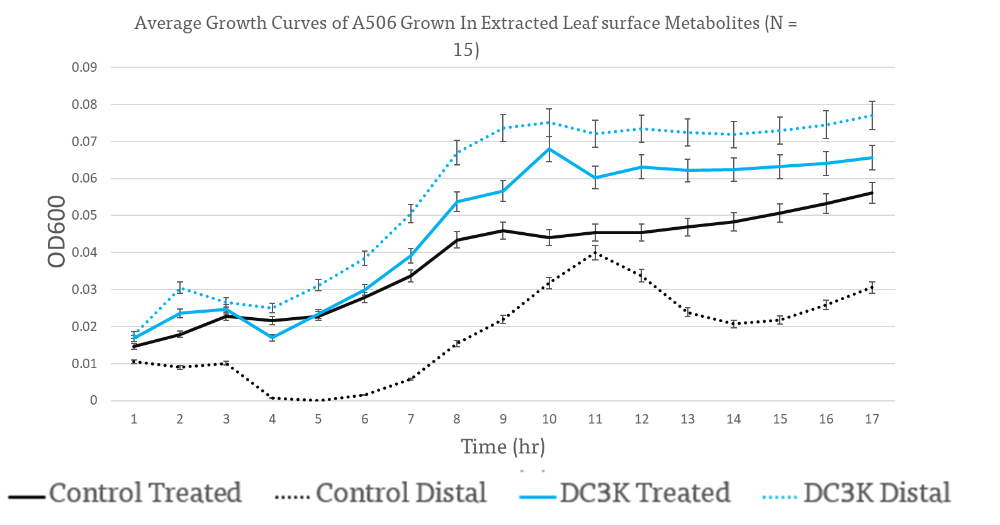
Leaf surface metabolites extracted from leaves distal to the inoculation with DC3K had metabolite profiles that significantly improved DC3K population size and growth (Figure 10).
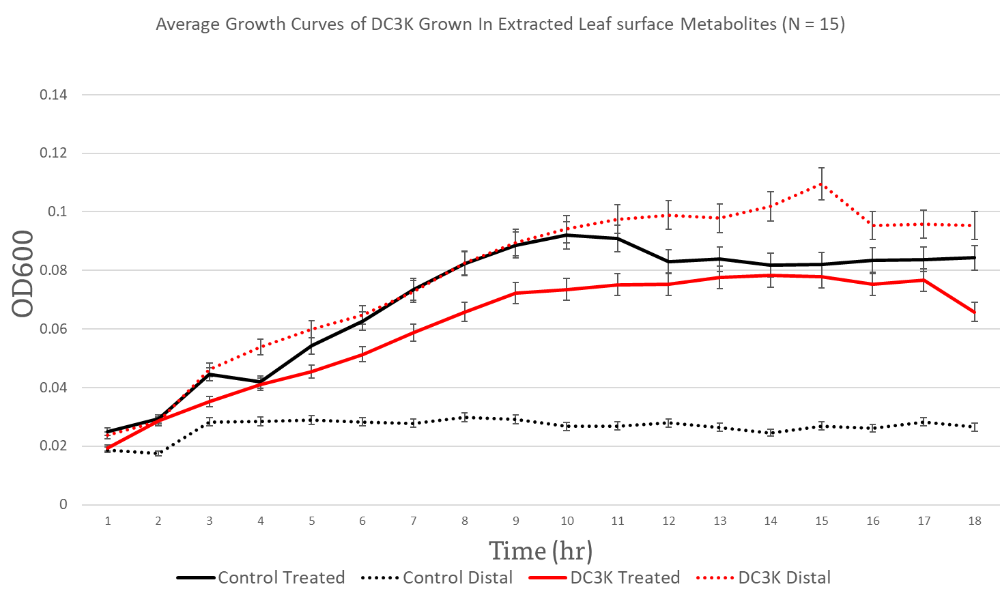
The control distal resulted in the lowest OD while the control resulted in a significantly higher growth rate than DC3K grown in distal LSMP or DC3K treated LSMP. A506 seemed to preferentially grow in leaf surface metabolite profiles extracted from distal leaves of DC3K treated plants, which supports a possible increase in sugar profile as observed in the HNMR. A506 reach similar OD highs as DC3K when grown in DC3K treated leaf surface metabolite profiles (Figure 11).
These were extremely interesting findings as it elaborates that DC3K may be altering the leaf surface metabolite to improve population growth and establishment, but other closely related organisms, or those with similar metabolic functions, may be able to utilize these compounds similarly.
Effects of leaf surface metabolite in vivo
Astoundingly, the results of this experiment indicated the plants infected with DC3K improve pathogen growth on subsequent leaves up to three (3) fold greater than populations established on uninfected plants (Figure 12).
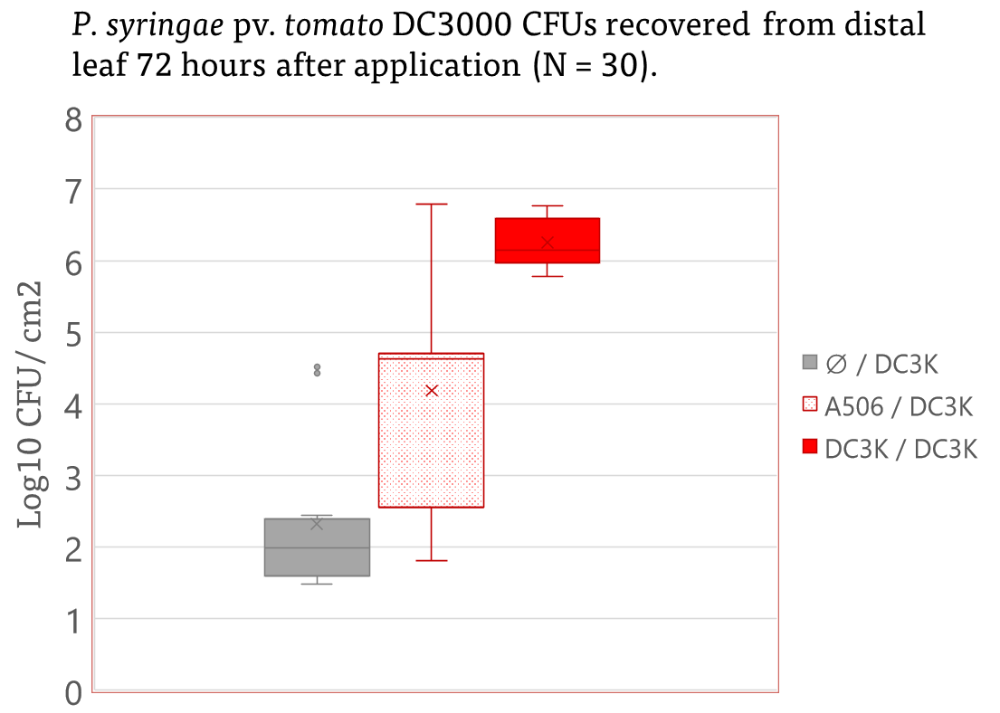
Furthermore, inoculated treated leaves with A506 and subsequent inoculation of the distal with DC3K resulted in 1-2 fold increase in recovered pathogen population. This draws parallels to other work in which recognition of a class of microbe associated molecular patterns can induce a specific response- it is possible that recognition of A506 elicits a host response that is beneficial to the pathogen. Further work must be done to elucidate this. No significant effect was identified on A506 populations when grown in DC3K infected plants (Figure 13).
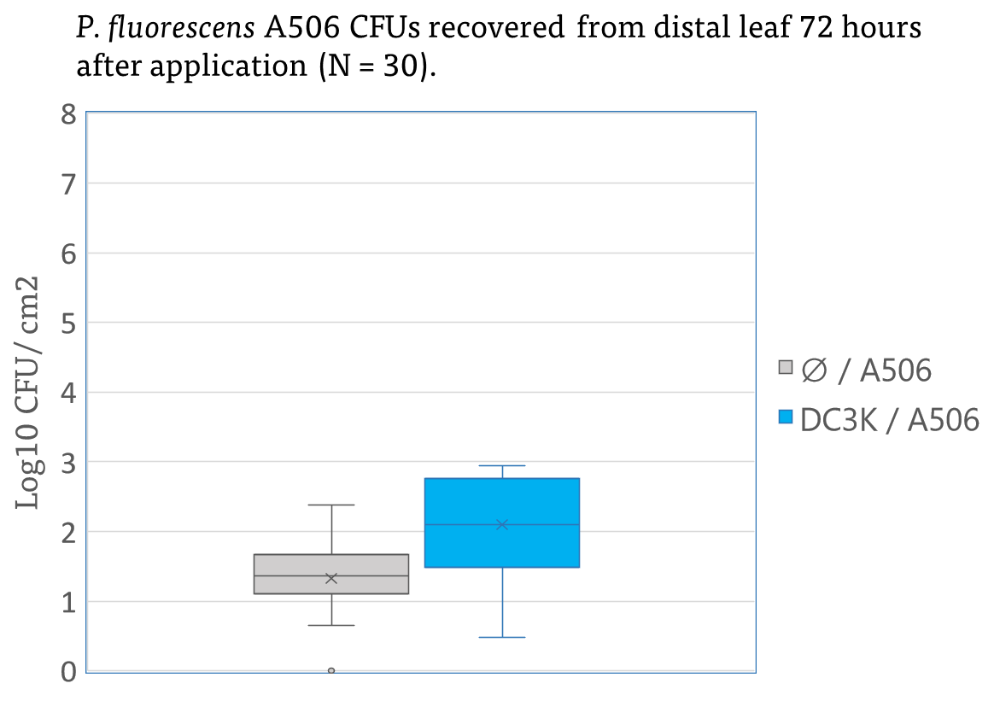
These results indicate a possible induced systemic susceptibility to DC3K once the pathogen is already present, which seems to be mediated in some part by the metabolites present on the leaf surface. Due to no difference in observable lesion amounts or disease symptoms, increased pathogen population has not been attributed to increased disease severity indicating that likely these metabolites changes aid in pathogen establishment.
In summary, the presence of a pathogen infecting leaves caused localized and systemic changes to leaf surface associated metabolite profiles (LSMP). These changes were predominantly associated with sugars and phenolics, and typically were increases in these compounds in distal leaves of DC3K infected plants. In non-host settings, the LSMP changes seem to improve the growth of both A506 and DC3K, but in a host setting DC3K LSMP manipulation fundamentally favors DC3K. While increased population establishment did not lead to visible increase in lesion amounts, this can not be discredited as the timeframe may have been too short for observing disease symptom differences. This indicates that once the pathogen is present within a system it may be extremely difficult if not impossible to eradicate disease.
Furthermore, this research highlights that not all plant growth promoters may be beneficial control agents for all diseases. A506 has often been seen as a robust plant growth promoter and heavily utilized across multiple systems, control of fire blight of pear or apple being the most notable. In this experiment, presence of A506 lead to a mild increase in pathogen population establishment. More work will need to be done to understand the extent of these induced systemic changes, but currently LSMP changes do lead to changes in bacterial population establishment which is a key factor in disease development.
Education & outreach activities and participation summary
Participation summary:
A major component of this work was to provide to the scientific community fundamental knowledge of how changes in leaf surface metabolites can affect associated bacteria, and how these changes are related to host defense responses. This is the first identified relationship between host defense response and external metabolite changes and their subsequent impact on bacterial assemblages. These results will support future in-depth studies identifying key biocontrol mechanisms and drivers of pathogen establishment for future breeding strategies.
This research has been presented within the US at the events below:
| Event | Location | Estimated Participants | Timing |
| Penn State One Health Microbiome Centre |
University Park, PA The Pennsylvania State University In-person and Zoom |
25 participants students/faculty/researchers |
March 2023 |
| Northeastern American Phytopathological Society Meeting |
Salisbury, CT Heritage Hotel In-person only |
50 participants students/faculty/researchers/public |
March 2023 |
| Penn State One Health Microbiome Centre |
University Park, PA The Pennsylvania State University In-person and Zoom |
100 participants students/faculty/researchers/public |
February 2024 |
Project Outcomes
This project highlighted the importance of host defense response in formation of bacterial assemblages via identifying mutability within LSMP and relating these changes to altered bacterial assemblage establishment and size. Altered leaf surface metabolite profiles in uninfected leaves of infected plants, or growth stimulated plants in case of A506, showcases how the changes in LSMP are a host-mediated response, as there is no other way to alter the LSMP. Improved pathogen growth in non-infected leaves in both A506 and DC3K infected plants indicate a host-mediated response that can be manipulated by members present on leaf assemblages. It also showcases the importance of assessing biocontrols within field settings across multiple stress signals. Biological agents may provide protection from one groups of pathogens but inevitably lead to susceptibility in others, which can lead to significant distrust in biologicals if utilized in fields settings inappropriately. However, this work showcases that the opposite can be true, that there may be biologicals capable of producing a LSMP that inhibits pathogen establishment but has not been identified yet. Continuing this work will not only lead to the divestment from synthetic pesticides but should hopefully improve biocontrol establishment and persistence in field settings, thus improving their adoption into normal agricultural practices.
This project increased my understanding of the skills required for, and the challenges involved with, metagenomic research that supports sustainable agriculture. Namely, I quickly realized it's very challenging to actively observe leaf surface metabolites without stripping the cuticle and often we would not get any metabolites after sample processing. Observing the differences between internal and external leaf (leaf surface) compartments is important as the interior is vastly more complex and has a higher concentration of all metabolites that will inevitably alter results if stripped- something else learned throughout this process. Furthermore, this work has highlighted the need to take these experiments into field settings for a better understanding of the biological relevance of this host-mediated effect. This immediately underlines the need for farmers to be present, not only for their valuable insights into phenotype development and pathogen severity, but to also have them buy into finding alternative approaches to conventional pesticides. This research also deeply highlights the need for integrated and multidisciplinary approaches towards answering questions regarding hot-pathogen interactions.
This project has generated new research ideas that I intend to develop as I progress into my career in host-pathogen mediated interactions. I've developed a keen understanding of metabolomic experiments as well as developed a novel process that allows me to identify differences in LSMP between a pathogen and a plant growth promoter via metabolic 'fingerprinting'. This opens up the doors to understanding these interactions across different pathosystems and crops, which I'm keenly interested in. Understanding trends in metabolites on leaf surfaces and their respective effect on microbial assemblages is a viable strategy for breeding, since secondary metabolites are a breeding trait and leaf surfaces metabolites are not overlapping with tomato fruit chemistry. I am interested to continue developing my research while involving more members of the agriculture community past researchers.