Final report for GNE21-255
Project Information
Maryland grain farmers are increasingly exploring organic production due to its potential economic and environmental benefits. However, the three-year transition period to organic certification presents considerable challenges, as farmers must adapt to non-chemical practices that limit their options for weed control, pest management, and nutrient supply. To ensure both profitability and environmental sustainability, it is crucial to identify practices that balance these factors. This project aimed to add a soil biological health component to the evaluation of organic transition strategies by examining microbial community composition, diversity, and functionality in soil. The goal was on understanding how different transition practices could promote soil health and sustainability.
The research study was conducted across multiple sites in Maryland, including both research stations and commercial farms, and compared four organic transition strategies: full but shallow (<12 cm) tillage with basic cover crops (Treatment 1), reduced tillage with mixed species cover crops (Treatment 2), minimal tillage and precision techniques with inter-seeded mixed cover crops (Treatment 3), and no tillage with perennial alfalfa-grass hay (Treatment 4). Microbial community including bacteria and fungi was assessed using advanced molecular techniques, such as DNA extraction, library preparation, and bioinformatics analysis. The diversity and composition of microbes were examined through alpha and beta diversity metrics, while microbial functionality was linked to nutrient cycling processes and fungal trophic modes.
The results highlighted that transition strategies significantly influenced microbial community diversity. Shannon diversity and Observed ASVs (Amplicon Sequence Variants - unique DNA sequences that represent specific types of microorganisms in a sample) increased significantly from 2020 to 2022 across all treatments, indicating rising microbial richness. However, Simpson diversity remained stable, suggesting that changes were driven by increased richness rather than shifts in community evenness. Fungal communities showed greater sensitivity to management practices, with perennial systems (T4) fostering distinct taxa adapted to low-disturbance environments. Reduced tillage and perennial systems (T2, T3, T4) supported higher microbial activity for nitrogen cycling pathways (e.g., nitrification, dissimilatory nitrate reduction) compared to full tillage (T1). Fungal trophic modes varied significantly, with perennial systems promoting saprotrophic (feeding dead and decaying matters) fungi but potentially accumulating pathogenic fungi. These findings emphasize the critical role that minimal tillage and perennial systems play in shaping microbial communities, which in turn enhances soil health, nutrient cycling, and carbon sequestration.
Although the study spanned only two years, it provides critical insights into how organic transition strategies influence soil microbial communities, which are key indicators of soil health and ecosystem services. The findings also emphasize the importance of site-specific management strategies, as local environmental conditions strongly mediate microbial responses. Reduced tillage and diverse cover crop systems emerged as promising practices for maintaining microbial diversity and functionality, while perennial systems showed potential trade-offs that warrant further investigation. Overall, the results highlight the potential benefits of adopting reduced tillage and cover cropping systems, not only in improving soil biological health but also in supporting sustainable farming practices that enhance soil fertility, ecosystem services, and environmental stability.
This project was part of a broader research program titled "Transitioning to organic grain production: strategies to increase viability and ecosystem services while reducing risks and obstacles," which was sponsored by the National Institute of Food and Agriculture (NIFA). The ultimate aim of the research project was to help boost the competitiveness of transitional organic grain crop producers by identifying system parameters that positively impact soil quality, crop yields, practicality and profitability. Microbiological indicators of soil health in the four transition systems were the focus of this NE-SARE project. The following were the specific objectives:
- Examine the effects of four organic transitional strategies on the composition and diversity of microbial communities.
The four systems were contrasted in terms of the types of prokaryotic communities (bacteria) and fungi, as well as their relative abundance and diversity. We hypothesized that the microbial community structure and diversity will differ among the systems and minimal tillage systems with higher biomass diverse cover crops will favor more microbial diversity.
- Study the effect of the transition strategies on potential microbial functionality.
We studied the relative abundances of enzyme encoding genes involved in certain functions and link that to potential microbial functionality (e.g. C degradation, N cycling) in the four systems by using databases such as Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway database for bacteria and UNITE database for fungi. We hypothesized that the functionality of microbes will differ between the treatments, such as: treatments with more legumes will favor nitrogen fixation pathways, perennial system will favor saprotrophic pathways, etc.
- Investigate the relationship between microbes and soil physical-chemical properties.
We investigated how microbial composition and individual microbial taxa relate to the soil physical and chemical properties.
Maryland grain farmers are showing increasing interest in organic production due to potential economic (much higher grain prices) and environmental (no synthetic pesticides) benefits. The three-year transitional period required for organic certification is a major barrier for prospective organic producers because they are limited in the methods allowable to control weeds, manage pests and diseases and supply plant nutrients (Delate and Cambardella, 2004). Particularly in Maryland, most conventional grain farms practice some form of no-till agriculture, grow cover crops and use strict nutrient management plans. This controlled nutrient, minimum tillage, cover cropping “eco-friendly” starting point in Maryland begs the question of whether transitioning to the organic farming practices from the “conventional” practices would cause an increase or decrease in environmental impacts and soil health. The answer most likely depends upon the system of practices (inputs, soil disturbance, soil cover, etc.). In Maryland, it is therefore essential to identify the organic transition strategies that can match or exceed the soil health and environmental benefits of the no-till with cover crops grain production system it would be replacing.
The purpose of this project was to add a biological component of soil health to the evaluation of four organic transition strategies done in the NIFA project. Specifically, we attempted to document the transition strategy (soil disturbance, soil cover, and input cost) that best contributed to the diversity and functionality of the microbial community in the soil system. To achieve these objectives, multi-farm experiments were conducted as a part of a larger NIFA grant project at four different sites in Maryland. Because the larger project (of which the NESARE project was a sub-study and enhancement) was designed to assess soil health impacts of the four transition strategies with a focus on soil physical and chemical properties, the NE-SARE grant was explicitly used to deepen the biological dimension of soil health investigation by assessing the microbial parameters using molecular techniques.
The strategies being compared and the microbial parameters proposed aimed to generate information to help farmers identify what management practices best contribute to the bio-indicators of soil health. Data on microbial diversity can infer nutrient cycling rates and defense against pathogens in a particular system (Fierer et al., 2020). Identifying microbial bio-indicators of several soil processes like nitrification, cellulose degradation, etc can be cheaper and easier alternatives to other direct methods (Fierer et al., 2020). Microbes respond rapidly to management practices, which can act as early indicators of changes in soil properties pertaining to the health of the environment (for example: abundance of a nitrifier taxa can indicate greater risks of losses of soil N via nitrification, leaching and potential groundwater pollution). Thus, the analysis of community structure, diversity and functionality of microbial taxa in the four systems is expected give a clearer picture of their relative productivity and sustainability and help in “fine-tuning” agronomic methods to the benefit of soil biology, overall environmental health and quality of agricultural systems.
Cooperators
- (Researcher)
Research
Treatments
We compared four organic grain transition strategies at various locations, including commercial and research farms. Three of the treatments involved grain cash crop production with different levels of soil disturbance, cover crop use, and input costs: full (but no plowing > 12 cm) tillage with basic cover crops (Treatment #1), reduced tillage with mixed species cover crops (Treatment #2), and precision techniques with inter-seeded mixed cover crops (Treatment #3). The fourth treatment (Treatment #4) employed perennial alfalfa-grass hay cultivation without any soil disturbance after initial establishment.
Study sites
In January 2020, field-scale plots were developed in four different locations in Maryland, including both university experiment station land (CMREC and LESREC) and commercial farms (Farm A also referred to as COOPER and Farm B). Three locations were on two different soil types that reflect the coarser and finer textured soils found in Delmarva's grain cropping coastal plain area; the fourth had loamy soils in the hilly Piedmont region. However, the commercial farm (Farm B) in the hilly Piedmont region was eliminated from the study later due to difficulty in application of treatments. In the commercial farm A, three transition systems (Trt #1, 2 and 3) were compared, and four systems were studied in experiment stations (Trt #1,2,3,4). The systems were tested in randomized complete block experiments, with the treatments repeated four times on each farm. The individual treatment plots range in size from 30 x 100 ft to 30 ft x 250 ft, large enough to allow commercial scale farm equipment to be used. In each treatment, two representative and relatively uniform sampling areas for collecting soil and plant samples were demarcated.
Sample collection
In early April 2020, the transition to organic farming commenced at the study sites with the planting of a uniformly managed spring oats crop, which was harvested in July 2020. During July and August 2020, composite soil samples were collected from each half of the treatment plots using 3.1 cm diameter × 30 cm long soil cores. The cores were subdivided into depth increments of 0–10 cm, 10–20 cm, and 20–30 cm. These soil samples represented the baseline soil conditions prior to the establishment of the four transition treatments. Baseline samples were analyzed for their physical and chemical properties, and subsamples of the 0–10 cm depth increment were stored unground and frozen at -20°C for soil microbial DNA extraction.
In the fall of 2022, soil samples were collected again as part of the NE-SARE-funded project. For each treatment, five cores were taken from each half of an experimental replicate and composited to create two composite samples per treatment. The cores were collected at least 2 m from the plot edges and distributed evenly throughout each plot. Debris was first removed from the soil surface, and a 3.1 cm diameter probe was inserted to a depth of 30 cm. The soil columns were then divided into 0–10 cm, 10–20 cm, and 20–30 cm depth increments, and samples from each depth were composited for each treatment.
To preserve the samples, the soil cores or loose soil were placed in sealed bags and immediately placed on ice. Between each composite sampling, gloves, probes, and knives were cleaned and sterilized with 70% isopropyl alcohol spray to prevent cross-contamination. Samples were transported to the laboratory in a cooler and frozen upon arrival. For microbial analyses, only the baseline soil samples (0–10 cm depth) collected in 2020 and the corresponding samples from 2022 (altogether 176 samples) were used.
Microbial analysis
Between December 2023 and August 2024, soil DNA extractions were conducted using homogenized, sieved (<2 mm), and thawed (previously stored at -20 °C) soil samples (250 mg). The extractions followed the QIAGEN DNeasy PowerSoil DNA extraction kit protocol (Qiagen, Hilden, Germany). The extracted DNA from each sample was quantified using a Qubit 2.0 fluorometer (Invitrogen, Waltham, MA, USA) and stored at -20 °C. When required, DNA concentrations were diluted to 5 ng/μl concentration for polymerase chain reaction (PCR) amplification of the extracted DNA segment.
For library preparation, the protocols outlined in the 16S Metagenomic Sequencing Library Preparation manual (Illumina, Part number 1504423 rev. B) were followed. To target prokaryotic communities (bacteria), a fragment of the 16S ribosomal RNA (rRNA) gene was amplified using primers 515F/806R, while fungal communities were analyzed by targeting the internal transcribed spacer (ITS) regions with primers ITS1f/ITS2r (Caporaso et al., 2011; White et al., 1990). Amplifications were performed using respective forward and reverse primers, 17.5 μl of Phusion Flash Master Mix (Thermo Fisher Scientific, Cat. No. F548S, Waltham, MA, USA), and 3.5 μl of DNA at 5 ng/μl.
Following amplification, two PCR cleanup steps were performed using AMPure XP beads (Beckman Coulter, Pasadena, CA, USA). Indexing (adding a unique sequence to each DNA template for identification) was conducted using the Nextera XT 96 Index Kit (Illumina, Cat. No. 15052165). The DNA was pooled, and the amplicon size of the library was assessed through high-sensitivity fluorometric quantification using the Qubit 2.0 fluorometer (Invitrogen, Waltham, MA, USA). The final libraries were sequenced on an Illumina MiSeq platform using a 600-cycle v3 cartridge at the University of Maryland Institute for Genome Sciences.
Illumina sequencing outputs in FASTQ format were processed using the dada2 package (v1.26.0; Callahan et al., 2016) in Rstudio Version: 2024.12.0+467. The sequences were quality-filtered, and adaptor and primer base pairs were removed. Error rates were then calculated, allowing paired-end reads to be merged. The dada2 package was further employed to remove chimeras (artifact sequences) and assign taxonomic classifications. Taxonomic assignments were made with reference to the SILVA database (v138.1, arb-silva.de) for prokaryotic 16S rRNA gene and the UNITE database (release 04.21.2024; version 10.0; Nilsson et al., 2019) for fungal ITS gene amplicon sequence variants (ASVs). The resulting ASV table was analyzed using the phyloseq (v3.20; McMurdie & Holmes, 2013) and vegan (v2.6-2; Oksanen et al., 2022) R package.
Before conducting microbial community analyses, features with ambiguous phylum-level annotations (classified as 'NA') and phyla with low prevalence were excluded. Additionally, low-abundance ASVs that were present in less than 1% of the samples were removed to ensure accurate and confident identification. To account for differences in sequencing depth across samples, rarefaction was performed by randomly subsampling to the minimum number of sequence reads present in a sample.
Statistical analysis
For Objective (1),
Alpha diversity measures were used to investigate how many different types of organisms live in a particular place, and how evenly those types are distributed. Alpha diversity metrics, including Shannon diversity (focusing on both the richness-the number of different species and evenness- how equally individuals are distributed among those species of a community), Simpson diversity (focusing more on the dominance of species within a community), and Observed ASVs (simple count of different microbial species detected in a sample), were calculated for both bacterial (16S rRNA) and fungal (ITS) datasets. Diversity estimates were computed from rarefied datasets using the phyloseq package in R. Boxplots were generated to visualize alpha diversity metrics across treatments and sampling years. Later, shannon diversity was used as the primary metric for exploring site and treatment effects. Additional visualizations included facet plots to investigate site-specific differences. The normality of diversity indices was tested using the Shapiro-Wilk test. For non-normal distributions, non-parametric statistical tests were applied. The Kruskal-Wallis test was used to evaluate overall differences in diversity among treatments, followed by Dunn’s post-hoc test for pairwise comparisons. Wilcoxon rank-sum tests were used to assess differences between sampling years for each treatment. P-values were adjusted using the Bonferroni correction to control for multiple testing. Site-level treatment effects were analyzed using separate Kruskal-Wallis tests for Shannon, Simpson, and Observed ASVs, followed by pairwise Wilcoxon tests for significant results. Significant results were annotated on the plots using statistical letters derived from post-hoc tests. Community composition for both bacterial and fungal taxa was analyzed by calculating relative abundances at the taxonomic levels of Class (16S rRNA) and Order (ITS). The ASV table was collapsed to the respective taxonomic levels using the tax_glom function from the phyloseq package, and relative abundances (proportion or percentage of each group in the total community) were calculated by normalizing read counts to the total number of sequences in each sample. Taxa with relative abundances below a threshold (1% for bacteria, 5% for fungi) were grouped into an "Other" category for simplicity and visualization clarity. Taxonomic data were aggregated by treatment, and mean relative abundances were computed for each taxonomic group. Stacked bar plots were created to visualize community composition across treatments, with relative abundances displayed as proportions.
Beta diversity, which tells how similar or different the microbial communities are between treatments over time and across sites was assessed to evaluate microbial community composition differences across treatments, years, and sites using Bray-Curtis dissimilarity. All analyses were performed separately for bacterial (16S rRNA) and fungal (ITS) communities using the phyloseq, vegan, and pairwiseAdonis packages in R. Bray-Curtis distance matrices were computed from the rarefied sequence dataset using the distance function in phyloseq. To test for significant differences in microbial community composition, analysis of similarity (ANOSIM) and permutational multivariate analysis of variance (PERMANOVA) were conducted. Pairwise PERMANOVA was conducted using the pairwise.adonis2 function to identify treatment pairs with statistically significant differences in microbial community composition.
For Objective (2),
Functional potential or functionality of microbes refers to the types of processes that microbial communities can potentially carry out, based on their genetic makeup. Instead of just counting the number or types of microbes, functionality looks at what these microbes can do—how they contribute to important soil processes like nutrient cycling, organic matter decomposition, and carbon sequestration. To infer the functional potential of the bacterial communities, Tax4Fun was employed, a computational tool that predicts KEGG Orthologs (KO) and functional pathways based on taxonomic profiles. The Tax4Fun pipeline was implemented using the themetagenomics package in R (Woloszynek et al., 2017). The SILVA reference database (silva_ko) was downloaded and used to match the 16S rRNA amplicon sequences to KEGG pathways. The relative gene abundances were calculated by summing all KEGG orthologs associated to a specific metabolic pathway (Cellulose degradation, N fixation, etc) (Lüneberg et al., 2018). Functional pathways were grouped into organic matter decomposition (cellulose, hemicellulose, lignin, and chitin breakdown), nitrogen cycling (nitrification, denitrification, nitrogen fixation, nitrate reduction, ammanox), and sulfur/methane metabolism (methane oxidation, sulfate reduction). KEGG pathway abundances were visualized using boxplots with significance letters indicating pairwise treatment differences. To characterize the ecological roles of fungal taxa, FUNGuild was used for trophic mode assignment. The FUNGuild analysis was performed using the FUNGuildR package in R. Fungal taxa were assigned to functional guilds using the funguild_assign() function. Taxa with confidence rankings of "Probable" and "Highly Probable" were retained for downstream analysis. Fungal communities were categorized into pathotrophic, saprotrophic, symbiotrophic, and mixed modes (e.g., pathotroph-saprotroph) trophic modes. Relative abundance was calculated for each trophic mode per sample. Normality of KEGG and trophic mode abundances across treatments and years (2020 vs. 2022) was assessed using the Shapiro-Wilk test. Since most distributions were non-normal, a Wilcoxon rank-sum test was conducted to compare functional pathway abundance between years within each treatment. For pathways showing significant differences, false discovery rate (FDR) correction was applied to account for multiple comparisons. To assess differences in abundance across treatments within 2022, a Kruskal-Wallis test was performed, followed by Dunn’s post-hoc test to identify significant pairwise treatment differences.
To investigate patterns or clusters of treatments or sites where microbial communities are more similar to each other, and to see how they differ across treatments or time, we used non-metric multidimensional scaling (NMDS). NMDS takes the Bray-Curtis dissimilarity (which measures how different the microbial communities are from each other) and tries to place the samples in a way that reflects those differences. For both datasets, raw sequencing reads were normalized to relative abundances, and NMDS ordinations were computed using the ordinate function with Bray-Curtis dissimilarity. For visualization, scatter plots of NMDS scores were generated using ggplot2, where points were color-coded by treatment and shaped according to site. For a 2022-specific NMDS ordination, samples from 2022 were subsetted, and NMDS ordinations were recomputed to assess treatment and site effects independently. To further explore the influence of environmental factors, soil property vectors were fitted onto the NMDS ordination using the envfit function from the vegan package. Soil parameters obtained from the NIFA study included bulk density, aggregate stability, permanganate-oxidizable carbon (POXC), and microbial respiration (mgC_kgsoil_24h and mgC_kgsoil_72h). Continuous soil variables were log-transformed before analysis to improve normality and reduce skewness. Only soil vectors that were significantly correlated with NMDS ordination axes (p < 0.05) were displayedin the NMDS plots. To identify the microbial taxa contributing most to variation in community structure, taxa were collapsed at the Class level for bacteria and Order level for fungi using the tax_glom function in phyloseq. Relative abundances were computed for each taxonomic group, and taxa contributing less than 1% of total relative abundance were filtered out to retain the most ecologically relevant groups. Significant taxonomic vectors were fitted onto the NMDS ordination using envfit, and taxa with p-values < 0.01 were plotted. The 20 most significant taxa were selected based on vector length, representing the strongest associations with microbial community variation.
Objective 1:
Alpha Diversity
Significant differences were observed in Shannon diversity and Observed ASVs between the two sampling years (2020 and 2022) across all treatments, but there were no significant differences in Simpson diversity. As shown in the Fig. 1, both Shannon diversity and Observed ASVs increased significantly from 2020 to 2022 across treatments, indicating an increase in microbial richness and diversity over time.
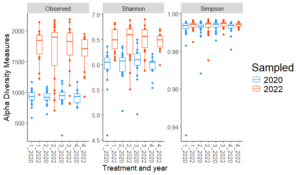
These temporal changes likely reflect cumulative effects of organic transitional strategies and environmental factors that gradually influenced microbial populations. The lack of significant differences in Simpson diversity between years suggests that while bacterial richness (Observed ASVs) and richness-evenness (Shannon diversity) increased, the overall evenness of bacterial communities (as captured by Simpson diversity) remained stable. This indicates that the temporal changes were driven primarily by an increase in richness rather than changes in the distribution of taxa abundances. With individual sites, no significant differences (p≤0.05) in any of the alpha diversity metrics were detected between treatments. This suggests that site-specific environmental factors, such as soil properties, climate, or historical management practices, have a stronger influence on alpha diversity than the transitional strategies applied.
For fungal communities, observed diversity (count of unique microbial species) showed significant differences between years in Treatments 1 and 2, with higher richness in 2022 compared to 2020 (Figure 2). This temporal increase suggests that fungal richness may respond positively to certain organic transition strategies over time. However, no significant differences in Shannon or Simpson diversity were observed between years within individual treatments, indicating that the evenness and diversity of fungal communities remained relatively stable over the study period.
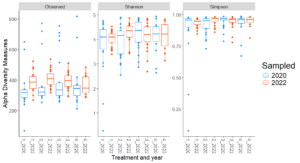
Site-level analyses revealed significant differences in diversity at CMREC and LESREC (Figure 3) in 2022. At CMREC, both Shannon and Simpson diversity metrics were significantly different across treatments. The Shannon diversity at LESREC also showed significant variation between treatments, while Simpson diversity remained consistent. In contrast, COOPER sites exhibited no significant differences in any diversity metrics across treatments or years, highlighting site-specific responses of fungal communities to organic management strategies. At CMRECC, Treatment 2 (Reduced tillage with multi-species moderate biomass cover crops) and Treatment 4 (Perennial alfalfa grass hay) exhibit significantly higher diversity compared to Treatment 1 (Full tillage with single-species cover crops). At LESREC, significant differences in shannon diversity were observed, with Treatment 4 showing the lowest diversity and differing significantly from Treatment 2, while Treatments 1 and 3 are statistically similar. The lower diversity observed under T4 at LESREC may be due to dominance by specific fungal taxa adapted to perennial systems, reducing overall evenness. This highlights the potential trade-offs in fungal diversity under perennial systems compared to systems with active cover crop rotations (T2).
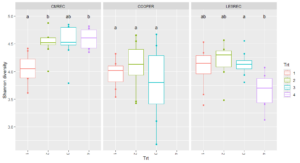
Beta Diversity
Beta diversity analyses, based on Bray-Curtis dissimilarity, revealed significant differences in bacterial community composition between the two years (2020 and 2022) within each treatment. ANOSIM results showed weak but statistically significant temporal differentiation, with R-statistics ranging from 0.2234 to 0.3471 (p ≤ 0.008) (Table 1).
Table 1: Analysis of similarities (ANOSIM) to test for differences in bacterial community composition between sampling years within each treatment.
|
Treatment |
ANOSIM R-Statistic |
p-value |
Interpretation |
|
Treatment 1 |
0.3152 |
0.001* |
Significant difference between years 2020 and 2022 |
|
Treatment 2 |
0.3209 |
0.001* |
Significant difference between years 2020 and 2022 |
|
Treatment 3 |
0.3471 |
0.001* |
Significant difference between years 2020 and 2022 |
|
Treatment 4 |
0.2234 |
0.008 |
Significant but weaker difference between years 2020 and 2022 |
These results suggest that bacterial communities have gradually diverged over time in each treamtment, potentially in response to organic management practices, changes in soil properties, or other environmental factors. In 2020 baseline samples, treatments did not influence microbial communities (p = 0.74, R = -0.01241). In 2022, treatments had a statisticall significant but very weak effect (p = 0.017, R = 0.04818). The weak but significant result in 2022 suggests that fungal communities could be slowly responding to treatment over time. Strong site-level differences in bacterial community composition were observed in both 2020 and 2022. ANOSIM results revealed highly significant differentiation between CMREC, COOPER, and LESREC, with R-statistics exceeding 0.7 (p < 0.001) in both years. Pairwise PERMANOVA confirmed significant site effects, with the greatest differentiation observed between CMREC and COOPER. These findings underscore the importance of site-specific environmental conditions, such as soil type, nutrient availability, and historical land use, in shaping bacterial community composition.
The ANOSIM results indicate significant but weak temporal shifts in fungal community composition across all treatments between 2020 and 2022 (Table 2). The R-statistic values ranged from 0.1171 (T1) to 0.2925 (T4), with the strongest shift observed in T4 (no-disturbance perennial systems). These findings suggest that fungal communities are gradually changing over time, likely influenced by the different management strategies employed. The more pronounced temporal changes in T4 may be attributed to the stability of the perennial system, which facilitates fungal succession and the establishment of fungal taxa adapted to less disturbed environments.
Table 2: ANOSIM results for fungal community composition across years 2020 and 2022 within each treatment)
|
Treatment |
ANOSIM R-Statistic |
p-value |
Interpretation |
|
Treatment 1 |
0.1171 |
0.002 |
Significant but weak difference |
|
Treatment 2 |
0.1663 |
0.001 |
Significant but weak difference |
|
Treatment 3 |
0.1744 |
0.001 |
Significant but weak difference |
|
Treatment 4 |
0.2925 |
0.001 |
Moderate difference |
ANOSIM results for 2022 revealed moderate-to-strong treatment effects at CMREC (R = 0.5216) and LESREC (R = 0.4219), while COOPER showed weak effects (R = 0.0937, p = 0.044) (Table 3).
Table 3: ANOSIM results for fungal community composition across treatments within each site (2022)
|
Site |
ANOSIM R-Statistic |
p-value |
Interpretation |
|
CMREC |
0.5216 |
0.001 |
Moderate-to-strong treatment effect |
|
LESREC |
0.4219 |
0.001 |
Moderate treatment effect |
|
COOPER |
0.0937 |
0.044 |
Weak but significant treatment effect |
Pairwise PERMANOVA results further highlighted the distinctiveness of T1 and T4, particularly at CMREC and LESREC (Table 4). For example, at CMREC, T1 vs. T4 had an R² of 0.2278, reflecting moderate treatment differentiation. LESREC exhibited similar patterns, with the strongest effect observed between T1 and T4 (R² = 0.2821). In all sites, T2 and T3 were not significantly different. These results suggest that fungal communities at CMREC and LESREC are more responsive to treatment-induced changes, possibly due to greater environmental variability or differences in soil management history. In contrast, the weaker effects at COOPER may indicate that other factors, such as inherent site stability, limiting the influence of treatments on fungal composition.
Table 4: Pairwise PERMANOVA Results for Fungal Community Composition Between Treatments Within Each Site (2022)
|
Site |
Comparison |
R² (Effect Size) |
p-value |
Interpretation |
|
CMREC |
T1 vs T2 |
0.2242 |
0.001 |
Moderate treatment effect |
|
CMREC |
T1 vs T3 |
0.1945 |
0.001 |
Moderate treatment effect |
|
CMREC |
T1 vs T4 |
0.2278 |
0.001 |
Moderate treatment effect |
|
CMREC |
T2 vs T3 |
0.0685 |
0.391 |
Not significant |
|
CMREC |
T2 vs T4 |
0.1574 |
0.001 |
Significant but weak effect |
|
CMREC |
T3 vs T4 |
0.1463 |
0.002 |
Significant but weak effect |
|
LESREC |
T1 vs T2 |
0.1116 |
0.041 |
Significant but weak effect |
|
LESREC |
T1 vs T3 |
0.1417 |
0.005 |
Moderate effect |
|
LESREC |
T1 vs T4 |
0.2821 |
0.001 |
Strong treatment effect |
|
LESREC |
T2 vs T3 |
0.0843 |
0.226 |
Not significant |
|
LESREC |
T2 vs T4 |
0.2468 |
0.001 |
Moderate treatment effect |
|
LESREC |
T3 vs T4 |
0.2092 |
0.001 |
Moderate treatment effect |
|
COOPER |
T1 vs T2 |
0.1098 |
0.032 |
Significant but weak effect |
|
COOPER |
T1 vs T3 |
0.0984 |
0.059 |
Marginally significant |
|
COOPER |
T2 vs T3 |
0.0696 |
0.382 |
Not significant |
The observed treatment effects, while weak overall, suggest that fungal communities are gradually responding to management practices, with the strongest changes associated with T1 (high soil disturbance) and T4 (low soil disturbance) and no changes in intermediate soil disturbance treatmens (T2 and T3). The distinctiveness of T1 and T4 highlights the impact of soil disturbance on fungal communities, with reduced disturbance favoring fungal taxa adapted to stable environments. The strong site differentiation highlights the importance of local environmental factors in shaping fungal community structure, which appears to be more sensitive to long-term conditions compared to bacterial communities.
Bacteria Venn Diagram (Figure 4) shows the shared and unique bacterial ASVs (Amplicon Sequence Variants) across the four treatments. In simple terms, each ASV is like a "barcode" that identifies a specific bacterial species or strain based on its genetic sequence. A significant overlap exists in the central portion, indicating a substantial core bacterial community (4453 shared ASVs) across all treatments. Treatment-specific regions (e.g., 1307 unique ASVs in T1) suggest that each treatment also fosters unique bacterial taxa. Treatments with reduced disturbance and diverse cover crops (T3, T4) show fewer unique ASVs, suggesting that these conditions promote more similar bacterial communities.
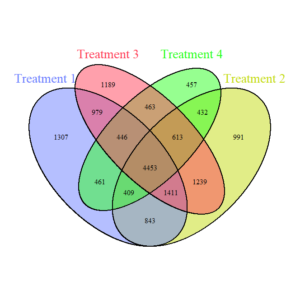
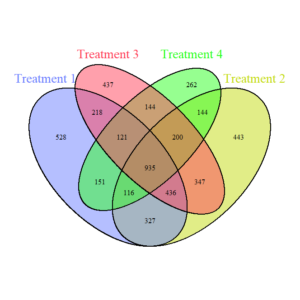
Fungi Venn Diagram (Figure 5) demonstrates the shared and unique fungal ASVs across treatments. The fungal core community (935 shared ASVs) is smaller compared to bacteria, reflecting greater fungal specificity in response to treatments. Unique ASVs are distributed across treatments, with T1 (full tillage) having the most unique ASVs (528), suggesting that high disturbance creates a distinct fungal niche.
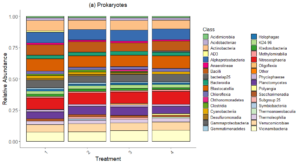
Bacteria Relative Abundance plot (Figure 6) highlights the distribution of bacterial classes across treatments in 2022 samples. Dominant classes include Actinobacteria, Bacteroidia, Alphaproteobacteria, Niotrososphaeria and Gammaproteobacteria, which remain relatively consistent across treatments. Minor differences in relative abundance suggest that bacterial composition is less sensitive to treatments compared to fungal communities.
Fungi Relative Abundance plot (Figure 7) displays fungal community composition at the order level for each treatment. Orders like Hypocreales, Pleosporales, and Agaricales, Mortierellales are dominant across treatments. T4 (perennial system) shows a distinct composition with more Pleosporales, Hypocereales, and Glomerellales, reflecting the effects of long-term no-till and perennial vegetation on fungal taxa. Treatments differ in the proportions of taxa associated with pathotrophic and saprotrophic functions, as evidenced in functional analysis later.

Both bacterial and fungal communities share a substantial core across treatments, but fungi exhibit greater specificity and sensitivity to treatment effects. The high number of unique ASVs in T1 for both bacteria and fungi suggests that intensive disturbance drives unique community assembly processes. The fungal community responds more strongly to reduced tillage and diverse cover crops (T3, T4), as evidenced by the relative abundance plots and the distinct taxa associated with perennial and no-disturbance systems. The bacterial community appears more stable across treatments, with minor shifts in dominant classes, likely due to their functional redundancy and resilience to management practices.
Objective 2:
Significant differences were observed in nitrification and dissimilatory nitrate reduction among the treatments (Figure 8). For dissimilatory nitrate reduction (nitrate use as a energy source), Treatment 1 exhibited significantly lower pathway abundances compared to Treatments 2 and numerically lower than treatments 3 and 4. The latter three treatments shared similar levels, indicating more comparable contributions to this pathway. Similarly, nitrification (conversion of ammonia to nitrate) showed a distinct trend where Treatment 2 exhibited higher abundance compared to Treatment 1 and 3, while Treatments 4 remained intermediate, suggesting some differences in the capacity for nitrification between the systems. No significant differences were observed for nitrogen fixation, denitrification, or anammox, indicating uniform contributions to these pathways across treatments. Similarly, no difference was found in the organic matter decomposition (Figure 9) and Sulfur and methane pathways (Figure 10).

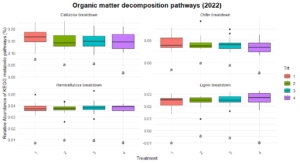
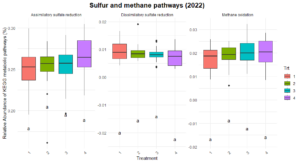
The differences observed in nitrification and dissimilatory nitrate reduction suggest a potential influence of tillage and vegetation type on the nitrogen cycling processes. Treatment 1, which represents full tillage with single-species cover crops, showed reduced contributions to dissimilatory nitrate reduction compared to Treatments 2, 3, and 4. This may reflect a disruption of microbial niches due to intensive tillage, resulting in lower microbial functional capacity for nitrate reduction. In contrast, the higher levels of dissimilatory nitrate reduction and nitrification in reduced tillage (Treatment 2) and perennial systems (Treatment 4) suggest more stable soil environments that support microbial activity likely due to increased substrate availability and reduced soil disturbance. The lack of significant differences in nitrogen fixation across treatments could indicate that this function is potentially constrained by environmental factors such as soil nutrient levels or moisture.
Significant differences in fungal trophic modes were evident for pathotroph and saprotroph abundances (Figure 11). Pathotroph abundance was significantly lower in Treatments 1 and 2 compared to Treatment 3, which had the highest abundance, while Treatment 4 exhibited intermediate levels. Saprotroph abundance was significantly higher in Treatment 4 compared to Treatments 1, 2, and 3, which were relatively similar. No significant differences were observed in other trophic modes, including symbiotrophic or mixed modes (Figure 12).
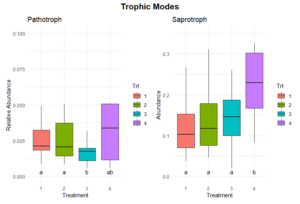
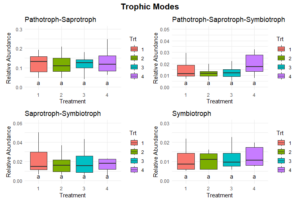
The significantly higher saprotroph abundance in Treatment 4 suggests that the perennial alfalfa grass hay system promotes fungal communities specialized in decomposing organic matter. This could be due to the continuous input of organic material and the absence of soil disturbance, which creates a stable habitat for these fungi. In contrast, the lower saprotrophic abundance in Treatments 1, 2, and 3 may reflect reduced organic matter availability or more frequent disruption of fungal networks due to tillage. The higher pathotroph abundance in Treatment 4 could indicate the accumulation of pathogenic fungi in the less disturbed or perennial system due to the lack of soil disturbance and crop rotation. This observation warrants further investigation into potential plant health risks associated with long-term perennial systems.
The results suggest that reduced and minimum tillage systems with diverse cover crops (Treatments 2 and 3) strike a balance between maintaining functional microbial processes (e.g., nitrogen cycling) and limiting potential negative effects such as the accumulation of pathogenic fungi. In contrast, the perennial system (Treatment 4) fosters a distinct microbial community that excels in organic matter decomposition but may require monitoring for pathotroph buildup. Treatment 1 (full tillage) consistently exhibited lower microbial functionality, highlighting the detrimental effects of intensive tillage on microbial processes.
Objective 3:
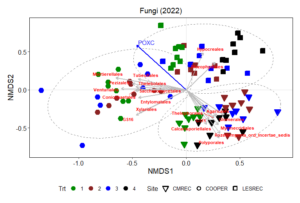
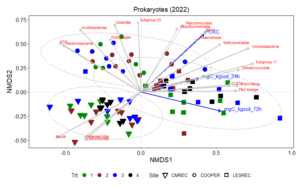
Non-metric multidimensional scaling (NMDS) plot results shows how microbial communities are distributed across different treatments and sites. The points represent microbial samples, and their position in the plot reflects similarity in community composition. Points that are close together indicate similar microbial communities, while points that are far apart indicate different communities. The NMDS ordinations for fungal (Figure 13) and prokaryotic communities (Figure 14) in 2022 reveal clear separation of microbial communities based on site and treatment within site. For fungi, the NMDS1 axis differentiates the treatments, while NMDS2 captures variability associated with site differences. Notably, treatments 3 and 4 (minimum tillage with high biomass cover crops and perennial no-tillage system, respectively) were more closely clustered, indicating higher similarity in fungal community composition. In contrast, treatments 1 and 2 (full tillage and reduced tillage systems) were more distinct, suggesting that increased disturbance disrupts fungal community structure. For prokaryotes, NMDS analysis reveals similar clustering patterns, with treatments 3 and 4 showing closeness and differentiation from treatments 1 and 2. The ordination plots also reveal site-level effects, with fungal and bacterial communities at each site exhibiting distinct divergence from each other. This differentiation aligns with soil physicochemical properties, which likely exert selective pressures on microbial community composition.
The NMDS results demonstrate that both fungal and bacterial community structures are significantly influenced by management practices and site-specific environmental conditions. The clustering of fungal communities and their association with higher POXC values suggest the development of functionally specialized fungal communities. Orders such as Thelephorales and Agaricales, which are known to form mutualistic relationships with plants, thrive under minimal disturbance, likely enhancing nutrient cycling and carbon sequestration. Prokaryotic communities, while also influenced by treatment, exhibited distinct responses. Classes such as Planctomycetes and Desulfomonadales were associated with POXC indicative of their role in organic matter decomposition and nutrient cycling. Soil respiration appears to be more pronounced in the LESREC site and specific bacterial taxa like Vicinamibacteria and Subgroup 11 seem to be linked with higher soil respiration, indicating that these bacteria might play a role in organic matter decomposition and carbon turnover.
The site-level differences observed for both fungi and bacteria highlight the importance of environmental and soil physicochemical properties in shaping microbial communities. These findings emphasize that site-specific management strategies are critical for optimizing soil health and microbial function. Moreover, the stronger association of POXC with both bacteria and fungi and soil respiration with the bacteria underscores their critical role in nutrient cycling and carbon stabilization.
This study shows the dynamic interplay between agricultural management practices, site-specific environmental conditions, and soil microbial community structure and function. Over the two-year study period (2020–2022), significant increases in Shannon diversity and Observed ASVs were observed, indicating an increase in microbial richness and diversity across all treatments. The temporal changes potentially reflect the cumulative effects of organic transitional strategies and environmental factors, which gradually shaped microbial populations. The stability of Simpson diversity suggests that the observed shifts were driven primarily due to increases in microbial richness rather than changes in community evenness. Site-related factors such as soil properties and historical management practices were stronger determinants of alpha diversity than the applied transitional strategies. This highlighted the importance of local environmental conditions in shaping microbial communities.
Fungal communities exhibited significant increases in richness over time, particularly in treatments with reduced tillage and diverse cover crops, highlighting their sensitivity to management practices. However, the stability of Shannon and Simpson diversity metrics indicates that fungal community evenness remained relatively consistent. Site-level analyses revealed significant differences in fungal diversity at CMREC and LESREC, with perennial systems (Treatment 4) promoting distinct fungal taxa adapted low-disturbance environments. In contrast, COOPER sites showed no significant treatment effects, emphasizing the role of inherent site stability in mediating fungal community responses.
Beta diversity analyses revealed significant temporal shifts in both bacterial and fungal communities, with weak but statistically significant differentiation between treatments. These shifts suggest gradual microbial community responses to organic management practices, soil property changes, and environmental factors. Strong site-level differentiation further established the role of local conditions, with bacterial communities showing greater resilience to treatment effects compared to fungi, which exhibited higher sensitivity to soil disturbance and vegetation type.
Functional analyses showed significant differences in nitrogen cycling pathways, with reduced tillage and perennial systems (Treatments 2, 3, and 4) supporting higher microbial activity for dissimilatory nitrate reduction and nitrification compared to full tillage (Treatment 1). Fungal trophic modes also varied significantly, with perennial systems promoting saprotrophic fungi specialized in organic matter decomposition but potentially accumulating pathogenic fungi, which requires further investigation into plant health risks.
These findings collectively emphasize the critical role of site-specific management strategies in optimizing soil health and microbial functionality. Reduced tillage and diverse cover crop systems strike a balance between maintaining microbial diversity and enhancing ecosystem services, while perennial systems may foster specialized communities with potential trade-offs. Our study underlines the importance of long-term monitoring and adaptive management to harness the benefits of microbial communities for sustainable agriculture. By integrating microbial community dynamics with soil health and crop performance, future research can refine management practices to enhance agricultural productivity, resilience, and environmental sustainability.
Education & outreach activities and participation summary
Participation summary:
We collaborated on the main NIFA project with commercial farmers, extension agents, and a non-profit farmer organization called Future Harvest which promotes sustainable agriculture in the Chesapeake bay region. As the NESARE grant covered important component of the larger research on soil health metrics, we integrated and will be including the ideas, methods, results, observations in the outreach activities pertaining to the larger project. Specific to this project, we discussed with farmers and extension agents at various stages of project implementation through in-person interactions, poster presentations and talks, which included details on what, why and how of our project, and our observations. An extension article titled "Interested in $10 corn and $30 soybeans for certified organic, but not sure how to transition?" was published in the November 2021 edition of University of Maryland Extension newsletter "Agronomy news". The article consisted of information on general introduction to the project and some preliminary results from the larger project.
I gave a oral presentation on "Strategies to Transition to Organic Grain: Impacts on Soil Health" in the ASA, CSSA, SSSA International Annual Meeting in Baltimore, MD in November, 2022. I presented about the impacts of organic transition strategies on the soil bulk density, labile carbon (Permanganate Oxidizable Carbon - POXC), and decomposition rate of standard tea material by the microbes. Similarly, a joint presentation on "Crop Productivity and Soil Health During Transition to Organic Farming" in a regional conference organized in January, 2023 by Future harvest, Chesapeake alliance for sustainable agriculture. We talked about the impacts of different organic transition strategies on the cover crop biomass, crop emergence, weed biomass, crop establishment, and yield. Similarly, we discussed the impacts on soil bulk density and permanganate oxidizable carbon (POXC).
I gave oral presentation on "Transitioning to Organic Grain Production in Maryland: Evaluating Soil-Health Impact of Four Strategies." in the ASA, CSSA, SSSA International Annual Meeting in St. Louis, Missouri in November, 2023. I presented a poster titled "Strategies to Transition to Organic Grain: Impacts on Soil Health and Crop Productivity" in University of Maryland, College of Agriculture and Natural Resources (AGNR) cornerstone event in October 2023 and November 2024. In those events, I presented about the impacts of organic transition strategies on several soil health parameters such as microbial respiration rate, aggregate stability, permanganate oxidizable carbon (POXC) and discussed with the attendees about the NESARE funded microbial study using DNA extraction and sequencing tools. Similarly, me and my advisor gave a join presentation on "Building Soil Health & Profitability while Transitioning to Organic Grains" in a regional conference organized in January, 2024 by Future harvest, Chesapeake alliance for sustainable agriculture. We talked about the impacts of different organic transition strategies on the soil health, cover crop biomass, crop emergence, weed biomass, crop establishment, yield, and profitability. On March 19, 2024, I presented on and interacted with farmers from Howard County, Maryland on the impacts of different organic transition strategies on soil health and crop productivity.
An assessment of the response of soil microbial communities to different organic transition strategies, along with recommendations as to the best strategy, will be the topic of at least one extension fact sheet or bulletin in 2025. The information from this project will be included in extension newsletters (i.e. UMD extension’s Agronomy news). Project results will also be reported at the meetings of professional societies (e.g., the American Society of Agronomy, Crop Science Society of America, and Soil Science Society of America tri-societies international annual meetings during 2025). The results from this project will also be published in a peer-reviewed journal (e.g., Agronomy Journal or Microbiology) with potential research paper titled "How the Transition to Organic Grain Effects Biological Indicators of Soil Health". The research paper will include our observations on the microbial diversity, composition and functionality from this study.
Details about selected talks/posters:
Gairhe, B., & Weil, R. (2024, May 8). Soil health and crop productivity impacts of four strategies for managing the
transition to organic grain. Presented at the Department of Environmental Science and Technology's Graduate Seminar,
University of Maryland College Park. https://www.youtube.com/watch?v=nW6F8vmHoGM&t=376s (>215 views)
Gairhe, B., & Weil, R. (2024, April 24). Transitioning to Organic Grain Production: Strategies to maximize profitability and
ecosystem services while reducing risks and barriers. Poster session presented at the USDA NIFA: OREI & ORG Project
Directors Meeting. Orlando, FL
Gairhe, B., & Weil, R. (2024, March 19). Building Soil Health and Profitable Production While Transitioning Conventional
Fields to Grow Organic Grain. Presented at the Howard County Mid Winter Ag Meeting, 2024.
Weil, R., & Gairhe, B. (2024, January 19). Building Soil Health & Profitability While Transitioning to Organic Grains.
Presented at the 2024 Annual Meeting of Future Harvest: Chesapeake Alliance for Sustainable Agriculture. College Park,
MD.
Gairhe, B., and Weil, R. (2023). Transitioning to organic grain production in Maryland: Evaluating soil-health impact of four
strategies. International annual meetings of the American Society of Agronomy and the Soil Science Society of America.
Indianapolis, IN. 30 Oct. 2023. ASA/CSSA/SSSA. https://scisoc.confex.com/scisoc/2023am/meetingapp.cgi/Paper/154129
Gairhe, B., & Weil, R. (2023, October 24). Strategies to Transition to Organic Grain: Impacts on Soil Health and Crop
Productivity. Poster session presented at the cornerstone event of the College of Agriculture and Natural Resources,
University of Maryland, College Park, MD.

Project Outcomes
In our project, we collaborated closely with 2 farm managers and a commercial farmer who played a pivotal role in implementing the different organic management treatments across the study sites. Additionally, we engaged with several farmers during presentations and workshops where we shared our insights and findings. Many of the farmers and farm managers interacted with us and reported changes in their knowledge and awareness about the significant role microbial communities play in soil health and agricultural productivity. They indicated a deeper understanding of how management strategies such as reduced tillage and diverse cover crops can influence microbial diversity, which is crucial for long-term soil fertility. Our collaboration also extended to undergraduate students, who were actively involved in key stages of the project, including soil sampling, sample processing, and DNA extraction. This hands-on experience helped to instill a deeper understanding of soil biology and its critical role in agricultural sustainability in the next generation of scientists and professionals. Once published, the technical data generated by this research will add valuable scientific knowledge to the fields of soil biology and organic farming, while also assisting farmers who are interested in adopting practices that enhance soil microbial health—particularly those that minimize soil disturbance and incorporate cover crops. Moreover, the project has strengthened the connection between researchers and farmers, empowering them with the knowledge and strategies needed to boost farm resilience and support long-term sustainability.
Throughout the course of this project, I gained a deeper understanding of the complex dynamics between agricultural practices and microbial communities, particularly in the context of transition to organic agriculture. Initially, we were focused on understanding how different management strategies, such as tillage and cover cropping, affected soil health and crop productivity. As I gathered further information and deeper study, it became evident that not only do these practices the physical and chemical aspects of soil health, but they also play a crucial role in shaping microbial communities responsible sustaining ecosystem services such as nutrient cycling, organic matter decomposition, and carbon sequestration. We became aware of the temporal and site-specific nature of microbial responses, emphasizing the importance of tailoring agricultural practices to local environmental conditions. This project also helped me develop several technical skills related to microbial research, including soil sampling for microbial studies, DNA extraction, library preparation, and bioinformatics data analysis. Through these hands-on experiences, I gained a deeper understanding of the complexity of microbial ecosystems and the challenges involved in studying them. I also learned that understanding the microbes in agricultural systems is essential but complex, requiring a balance of various factors. Sustainable agriculture must strike a balance between soil health, crop productivity, and profitability to ensure long-term success. Moving forward, I aim to explore further the role of microbes in sustainable agriculture, with a particular focus on the carbon cycle. This area holds great potential for improving soil health and mitigating climate change, and I am eager to continue investigating how microbial communities contribute to these processes in sustainable agricultural systems.